A novel biomarker measured early and able to predict future performance is an ideal strategy for the selection of high-performing sows.
June 2, 2020

Gilt development is crucial for the success of commercial sow farms. During acclimation, F1 gilts are evaluated for multiple factors with the objective of identifying those that will be efficient mothers with a high number of healthy piglets and many parities. In humans, the vaginal microbiota, which represents the collective bacteria present in the vagina, has been associated with reproductive outcomes (Choi and Blaster, 2012). However, this novel area of research has not been vastly explored in the reproductive pig. Hence, there is opportunity to identify novel biomarkers that can be used to predict farrowing performance.
Last January, we described how the pig genome partially controls the abundance of hundreds of bacteria in the vaginal tissue of F1 gilts. Further, in February we demonstrated how these bacteria are highly predictive of contrasting farrowing outcomes in F1 gilts.
Prior to farrowing, 302 F1 gilts from the same commercial sow farm had swab collected for identification and quantification of the microbiota in the vaginal tissue of each animal. The data are expressed as operational taxonomic units, which represent bacteria that are clustered together based on sequencing similarities. Collection of vaginal swabs occurred on Day 4 and Day 52 after porcine reproductive and respiratory syndrome modified-live vaccine vaccination. Subsequently, litter size traits were recorded on all animals. All F1 gilts had genotype data including over 50,000 genetic markers.
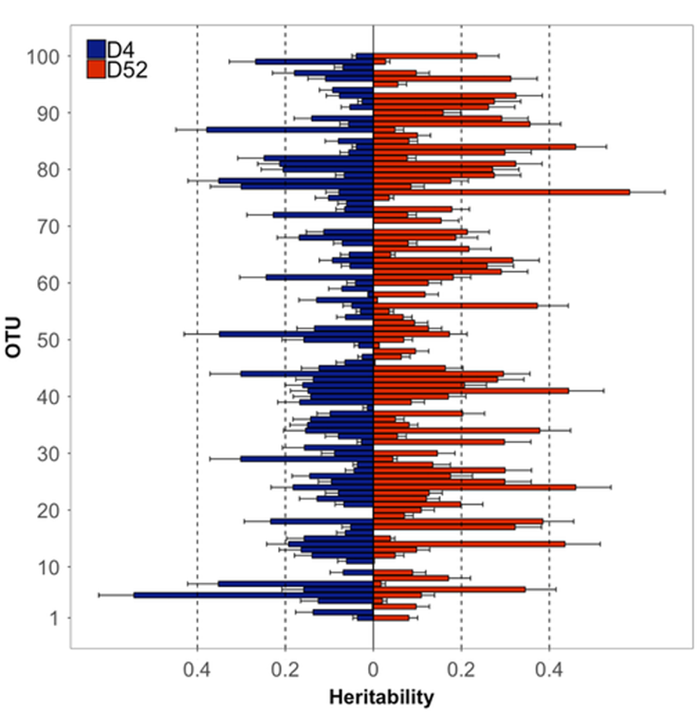
Overall, more than 2,000 OTUs were identified in the vaginal microbiota of commercial F1 gilts. The five most abundant phyla were Fusobacteria, Proteobacteria, Firmicutes, Bacteroidetes and Tenericutes. We estimated the gilt's genetics contribution (i.e. heritability) for the 100 most abundant OTUs (Figure 1). Ranging from the most (OTU 1) to the least (OTU 100) abundant OTU, we observed that heritability estimates ranged from low (below 0.1) to high (over 0.4) on both days of collection. In general, heritabilities were greater on Day 52 compared to Day 4. These results indicate that the vaginal microbiota of F1 gilts is impacted by the gilt's genetics and that genetic selection for changes in the abundance of specific bacteria is possible.
From our genomic analyses, we identified 22 regions in the pig genome highly associated (P<0.00001) with OTU relative abundance. Of these results, one region on chromosome 12 of the pig was identified for four OTUs. Several genes with a wide range of functions are located in this region, limiting our ability to identify a potential candidate gene controlling the abundance of these OTUs. Nonetheless, our results suggest that major regions in the gilt's genome control the relative abundance of OTUs, indicating that genetic markers could be used to efficiently genetically change the vaginal microbiota of gilts.
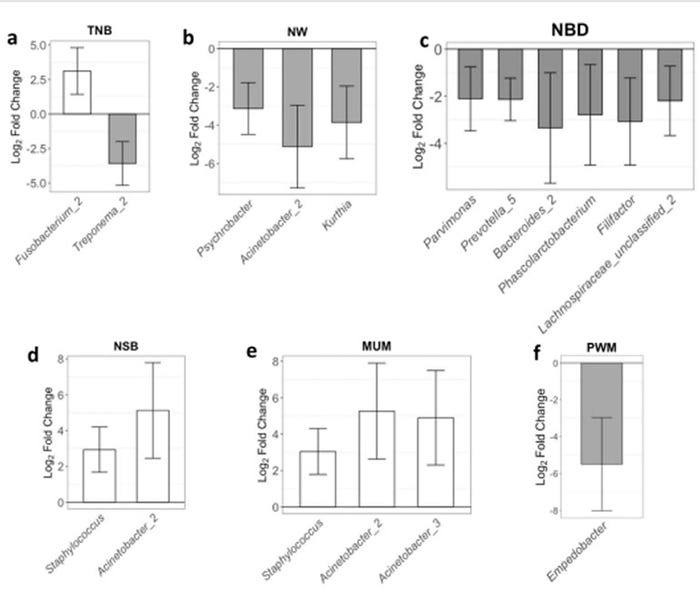
Finally, first-parity performance of these animals was used to identify the OTUs that could be used as biomarkers for improved performance. At first-parity, animals were classified as either having high or low performance (10 sows in each group) based on each farrowing trait: total number born, number born alive, number weaned, number born dead, number of stillborn, number born mummified and pre-weaning mortality. We identified (Padjusted<0.05) 13 different OTUs associated with TNB, NW, NBD, NSB, MUM and PWM (Figure 2). Most of the OTUs associated with contrasting farrowing performance had greater abundance in the gilts belonging to the low-performance group.
In addition, we predicted future farrowing performance (high or low) based on microbial data and obtained 100% of correct classification of gilts into their future farrowing performance group. Therefore, the vaginal microbiota collected prior to insemination showed to be highly predictive of first-parity performance.
Results from these studies indicate that the vaginal microbiota of gilts has the potential to be used to predict future farrowing performance. Thus, bacteria associated with higher reproductive performance could be identified and animals could be genetically selected to have increased abundance of "good" bacteria. The use of a novel biomarker that can be measured earlier in life and is able to predict future performance is an ideal strategy for the selection of high-performing sows, increasing the productivity and efficiency of the swine industry.
The authors would like to Smithfield Premium Genetics for their continued help and support. Questions can be directed to Nick Serão.
References
Cho I, MJ Blaser. The human microbiome: at the interface of health and disease. Nature Reviews Genetics. 13, 260-70 (2012).
Sanglard LP, S Schmitz-Esser, KA Gray, DCL Linhares, CJ Yeoman, JCM Dekkers, MC Niederwerder, NVL Serão. Investigating the relationship between vaginal microbiome and host-genetics and their impact on immune response and farrowing traits in commercial gilts. Journal of Animal Breeding and Genetics, 137(1):84-102 (2020).
Sanglard LP, S Schmitz-Esser, KA Gray, DCL Linhares, CJ Yeoman, JCM Dekkers, MC Niederwerder, NVL Serão. Vaginal microbiota diverges in sows with low and high reproductive performance after porcine reproductive and respiratory syndrome vaccination. Scientific Reports, 10:3046 (2020).
Sources: Nick Serão and Leticia Sanglard, who are solely responsible for the information provided, and wholly own the information. Informa Business Media and all its subsidiaries are not responsible for any of the content contained in this information asset.
You May Also Like



