Blueprint: Identification of the serotype-specific proteins in African swine fever
Facilitates vaccine design and development
April 11, 2016

African swine fever is an emerging disease threat for the swine industry worldwide. No ASF vaccine is available, and progress is hindered by lack of knowledge concerning the extent of ASFV strain diversity and the viral antigens responsible for protection in the pig. Available data from vaccination/challenge experiments in pigs indicate ASF protective immunity may be hemadsorption inhibition serotype-specific.
A better understanding of ASFV HAI serologic groups and their diversity in nature, as well as improved methods to serotype ASFV isolates, is needed. Here, we demonstrate that the genetic locus encoding ASFV CD2v and C-type lectin proteins mediates HAI serologic specificity and that CD2v/C-type lectin genotyping provides a simple method to group ASF viruses by serotype, thus facilitating study of ASFV strain diversity in nature and providing information necessary for eventual vaccine design, development and efficacious use.
ASF is a highly contagious, acute, viral hemorrhagic disease in domestic swine with mortality rates approaching 100%. There is no vaccine for ASF, thus leaving animal slaughter the only effective disease control strategy. Ongoing and devastating ASF disease outbreaks in the Caucasus region and Russia (2007 to present) highlight the significance of this disease threat for domestic swine populations. Starting from a single introduction in Georgia, ASF spread widely in the Caucasus and thousands of miles northward into Russia, reaching the Baltics, Poland and Ukraine, and now threatening at the doorstep of western Europe. The current endemic disease situation in Russia and the Caucasus, and the notable lack of a vaccine, illustrates the risk of ASF incursions into swine populations worldwide. Clearly, a similar epizootic in North America would be devastating to its swine industry. Many factors, including natural reservoirs of ASF in Africa (and now in Russia and the Caucasus), potential for establishment of an endemic ASF in new regions, rapid and efficient transmission of ASF among pigs, and relative stability of the virus in the environment, heighten the ASF threat and present challenges for its control.
Development of improved disease control strategies has been hindered by large gaps in knowledge concerning the causal agent, ASFV, and by the complexity of virus-host interactions underlying infection and immunity. Although there is no ASF vaccine available, it is clear that vaccination is possible, since protection against disease has been demonstrated using experimental, live-attenuated virus vaccines.
Unfortunately, natural antigenic diversity among ASFV strains limits utility of any vaccine based on a single strain. In addition, issues of residual pathogenicity and long-term viral persistence associated with current live-attenuated virus vaccine candidates render them unsuitable for use in nonendemic regions. Development of ASF subunit or vectored vaccines is slowed by lack of knowledge concerning the viral antigens responsible for inducing protective immunity and the extent of ASFV strain variation (Tulman et al., 2009). Progress on an ASF vaccine will require addressing these critical knowledge gaps.
Unlike many viruses currently challenging the swine industry, such as porcine reproductive and respiratory syndrome virus, ASFV is a DNA virus and thus much less variable over time. Once protective antigens have been identified, and the diversity of viral strains currently circulating in natural reservoirs determined, vaccines effective against relevant ASFV strains could be developed. Ideally, a vaccine would be not only efficacious, but also differentiate infected from vaccinated animal compatible, and thus suitable for use under emergency conditions in nonendemic countries of North America and western Europe.
ASFV, classified as the sole member of a new DNA virus family, Asfarviridae (Asfar, African swine fever and related viruses), is a large, genetically complex virus containing a double-stranded DNA genome of approximately 190 kilobase pairs that encodes more than 170 proteins. Comparative genomic sequencing of virulent field isolates and attenuated, cell-culture-adapted strains have identified terminal genomic regions as being variable in gene content and similarity, but also select genes throughout the genome that exhibit exceptionally high levels of sequence variation between viral isolates (Figure 1). While viral genomics is beginning to indicate candidate gene targets responsible for virulence and strain-specific protective immunity, to date, no ASFV proteins (antigens) have been definitely associated with solid protective immune responses in the pig (Tulman et al., 2009). Before ASF vaccine strategies can be designed and delivery systems evaluated, relevant protective antigens and the breadth of their natural diversity need to be known.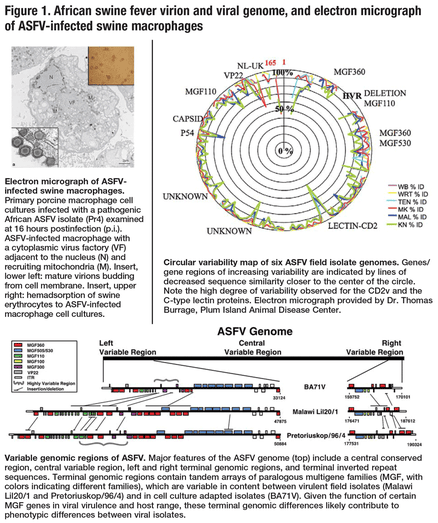
Although ASFV serologic assays used for disease diagnosis have focused on conserved, cross-reactive viral proteins suitable for detecting any viral strain, evidence indicates that distinct antigenic types of ASFV are discernable based on hemadsorption inhibition (HAI) serologic typing. Eight ASFV serogroups have been identified, although more likely exist. Notably and of particular significance for vaccine development, ASF protective immunity appears to be HAI serotype-specific, as viruses within a serogroup cross protect against one another. Furthermore, we have demonstrated HAI typing places ASFV into serogroups not necessarily resolved by conventional ASFV genotyping based on the capsid protein (P72) gene, as ASFV of HAI serogroups 1, 2 and 4 fall into the single P72 genotype I (Malogolovkin et al., 2015a).
Understanding of ASFV HAI serogroups, the viral determinants defining them and their diversity in nature may be of considerable significance for vaccine development. Two viral proteins encoded in the ASFV genome, CD2v (EP402R) and C-type lectin (EP153R), have been shown necessary and/or sufficient for ASFV hemadsorption (HA) in vitro. Also, comparative viral genomics have indicated that the locus containing adjacent CD2v and C-type lectin genes is among the most variable in the ASFV genome (Tulman et al., 2009). Thus, ASFV CD2v and C-type lectin are candidate proteins for mediating HAI.
Characterization of the ASFV CD2v and C-type lectin genes
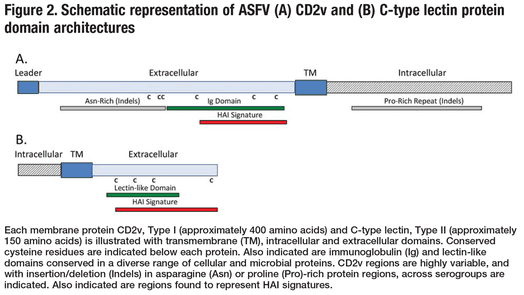
To examine the association of viral CD2v and C-type lectin proteins with ASFV serogroup specificity, we conducted a comparative analysis of these genes from a collection of ASFV isolates which had previously been characterized as to serogroup. Data from diverse viral isolates supported high levels sequence variation in previously described intra- and extracellular domains of the CD2v type I membrane protein and C-lectin type II membrane protein (Figure 2), consistent with previous reports of increased variability at the lectin/CD2v locus in the ASFV genome (Chapman et al., 2008; Tulman et al., 2009). CD2v sequence variation was observed in repeat domains, including the cytoplasmic proline-rich repeat domain which mediates interaction of CD2v with cellular protein(s). Variability in extracellular domains was noted as involving regions rich in asparagine (Figure 2), and asparagine variation in CD2v and C-type lectin impacted serogroup-specific differences in predicted N-glycosylation (17-25 and 7-9 total potential N-glycosylation sites, respectively). Variation in protein glycosylation has great potential to affect ligand-binding or antigenic specificities of CD2v and C-type-lectin glycoproteins. Conceivably, these differences in CD2v and C-type lectin primary sequence affect or define the structure and/or glycosylation potential important for hemadsorption and, moreover, the specificity which defines the HAI phenotype and ASFV cross protection. Elucidation of CD2v and C-type lectin glyco-variation and its impact on protective phenotypes is needed.
CD2v and C-type lectin-based phylogenetic groupings support a role for one or both proteins mediating HAI serospecificity
Given CD2v and C-type lectin variability and their importance in HA serogrouping, and notably given lack of serospecific resolution using standard P72 capsid gene typing (Malogolovkin et al., 2015), a correlation between CD2v/C-type lectin genotype and HAI phenotype was examined. Suitable extracellular protein subdomains (Figure 2) were selected for phylogenetic analyses, and data indicated that both CD2v and C-type lectin-based phylogenetic groupings correlated with serogroup (Figure 3). Sequences from viruses of similar serotype were often identical or nearly identical, grouping closely and reliably within the tree relative to isolates of different serotype. CD2v and C-type lectin protein domains thus represent signatures for ASFV serospecificity.
Observed through CD2v/C-type lectin HAI signature genotyping were ASFV isolates not clustering with, and with increased genetic distance to, known serospecific groups. Most notable among these were four isolates (Magadi, Davis, Bartlett, Killearn) previously untypable with SG1-SG8 reference antisera. Genetic outliers to reference serogroups, highlighted by those lacking reference-serum HAI reactivity, potentially represent viruses of novel, and yet-to-be-established serogroups. Identification of ASFV isolates outside of characterized serogroups is critical in considering all potential antigenic types for potential vaccine strategies of the future.
ASFV CD2v/lectin proteins mediate HAI serologic specificity
To assess the direct role of CD2v and C-type lectin proteins in ASFV HAI serospecificity, chimeric recombinant ASFVs in which CD2v/C-type lectin genes were exchanged between ASFV strains were constructed. Parental viruses were used to generate chimeric viruses in which native CD2v/C-type lectin genes were deleted and replaced by those from the virus of a different serogroup. While parental viruses maintained their respective serospecificity, HAI serogroup specificity was shifted in the chimeric viruses. A serogroup 2 chimeric virus (containing the CD2v/C-type lectin genes from the serogroup 4 virus) now exhibited serogroup 4 specificity instead of serogroup 2 specificity. Likewise, the serogroup 4 chimeric virus (containing the CD2v/C-type lectin genes from the SG2 virus) exhibited serogroup 2 specificity but not serogroup 4 specificity (Table 1). Thus, ASFV CD2v and/or C-type lectin proteins are necessary and sufficient for mediating HAI serologic specificity.
Overall, our data indicate that ASFV CD2v and/or C-type lectin proteins are necessary and sufficient for mediating HAI serologic specificity and that ASFV CD2v and C-type lectin signature sequencing provides a simplified method to group ASF viruses by serotype. CD2v/C-type lectin genotyping will facilitate study of ASFV strain diversity and antigenic variability and provide information critical for vaccine design, development and use.
Acknowledgements
This work was supported by the National Pork Board (grant 12-106), the USDA National Institute of Food and Agriculture (grant 2013-67015-21335) and the Russian Science Foundation (grant 16-16-00090).
You May Also Like



