Mapping hotspots for spread of PRRSV-2 lineage 1 in the United States
Finding contrasts to past study conclusions on PRRSV2- spread, which showed that geographical hub of PRRSV-2 L1 emergence was limited to the Midwest.
August 9, 2022
Porcine reproductive and respiratory syndrome costs the United States over $500 million annually (1).The repeated emergence of new genetic variants of PRRSV-2, the causative virus, reflects the failure of previous control efforts.
Within the United States, PRRSV-2 persistence is characterized by the co-circulation of numerous distinct variants defined by genetic variation of open reading frame 5 (ORF5), the major viral envelope glycoprotein coding gene. Although it is hypothesized that immune-mediated selection underpins its evolutionary dynamics, understanding spatial variability in selection pressures, variant emergence and spread is critical.
We investigated how the pace of evolutionary change varies across time and space, identified geographic hotspots for emergence of new variants and mapped patterns of inter-regional spread of PRRSV-2 Lineage 1 (L1), which has been the dominant phylogenetic lineage in the United States for the past 10 years (2). This lineage has diverged into at least seven sub-lineages (1A to H).
Utilizing representative subsets drawn from >40,000 ORF5 sequences, phylogeographic models were constructed to infer the ancestral dates of emergence for each L1 sub-lineage, frequency of historical spreading events between regions and viral population dynamics.
The most recent common ancestor of PRRSV-2 L1 in the United States was around 1986 (range: 1982-1988) in Canada (100% confidence). The virus population size in the United States displays cyclic peaks every ~6 years, on average. Sub-lineages 1C and 1A-2 mainly contributed to the recent peaks within the past decade (2011-2020).
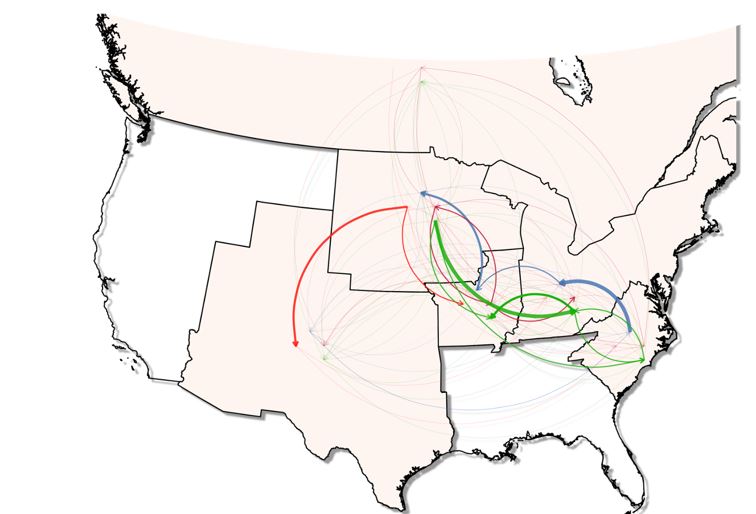
We also show that the Upper Midwest region was a hub for early L1 transmission (such as lineage L1F), likely because L1 viruses were introduced there from Canada and subsequently transmitted to other parts of the United States. The Upper Midwest and the East were the origin of sub-lineages 1C (emerging in 1995-2000) and 1A-2 (emerging in 2008-2012), respectively (Fig.1). This finding contrasts to the conclusion from past studies on PRRSV2- spread conducted in earlier decades, which showed that geographical hub of PRRSV-2 L1 emergence was limited to the Midwestern United States (5).
Such results reflect new or altered epidemiological dynamics caused by the evolution of co-existing PRRSV-2 variants. Altogether, our study provides an update on PRRSV-2 circulation pathways across the country, which could help strategize disease control and containment of emerging variants.
Literatures cited
1. Holtkamp DJ, Kliebenstein JB, Neumann EJ, Zimmerman JJ, Rotto HF, Yoder TK, et al. Assessment of the economic impact of porcine reproductive and respiratory syndrome virus on United States pork producers. J Swine Heal Prod. 2013;21:72–84.
2. Paploski IAD, Pamornchainavakul N, Makau DN, Rovira A, Corzo CA, Schroeder DC, et al. Phylogenetic structure and sequential dominance of sub-lineages of prrsv type-2 lineage 1 in the United States. Vaccines. 2021;9(6),608.
3. Suchard MA, Lemey P, Baele G, Ayres DL, Drummond AJ, Rambaut A. Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol. 2018;4(1):vey016.
4. Hadfield J, Megill C, Bell SM, Huddleston J, Potter B, Callender C, et al. NextStrain: Real-time tracking of pathogen evolution. Bioinformatics. 2018;34(23):4121–3.
5. Shi M, Lemey P, Singh Brar M, Suchard MA, Murtaugh MP, Carman S, et al. The spread of type 2 porcine reproductive and respiratory syndrome virus (prrsv) in North America: A phylogeographic approach. Virology. 2013.
Source: Nakarin Pamornchainavakul, Igor AD Paploski, Dennis N Makau, Mariana Kikuti, Albert Rovira, Cesar A Corzo, Kimberly VanderWaal and Samantha Lycett, who are solely responsible for the information provided, and wholly own the information. Informa Business Media and all its subsidiaries are not responsible for any of the content contained in this information asset.
You May Also Like



