The number of pigs received into neighboring farms was an important predictor of porcine epidemic diarrhea virus infection risk.
August 8, 2017

By Gustavo Machado, Andres Perez and Kim VanderWaal, University of Minnesota College of Veterinary Medicine Department of Veterinary Population Medicine
The movement of live pigs between farms is an important mechanism for disease introduction and spread. Thus, understanding the structure of livestock contacts and studying the routes, volumes, frequency and the risks associated with animal movement is a prerequisite for effective disease surveillance and control in animal populations1. At the same time, local area spread between neighboring farms is also implicated in the spread of viruses such as porcine epidemic diarrhea virus and porcine reproductive and respiratory syndrome virus.
The relative importance of these different diseases transmission mechanisms in determining between-farm spread in swine still uncertain and under debate2, which has implications for decision making in regards to appropriate biosecurity/surveillance measures. Much of our understanding of between-farm transmission in U.S. swine comes from outbreak investigations and case reports involving a relatively small number of farms. Large-scale datasets in which to test the relative roles of animal movements versus local area spread in between-farm spread are largely lacking.
Previous studies have used movement data to conduct network analyses describing contact between/among farms, slaughterhouses and regions. However, network analyses usually neglect alternative factors that affect the nature of direct and indirect contact between farms, such as environmental factors and geographic distance, which both may relate to local areas spread. To gain a more complete picture, we need to jointly address environmental/landscape factors, pig movements and spatial relationships among neighborhoods of swine farms, all of which likely interact to create between-farm transmission opportunities.
Collaborating with the Morrison Swine Health Monitoring Project, we analyzed the risk of sow farms breaking with PED based on the likelihood of exposure from neighboring farms, accounting for “neighborhood effects” such as pig movements into the neighboring farms and environmental factors that could affect viral dissemination and survivability. Determining the relative contribution of neighborhood effects, including pig movements, local dispersion and environmental/landscape factors, advances our understanding of the epidemiology of between-farm disease spread.
In order to understand the association between PED breaks in sow farms and possible predicting variables, we used a series of machine-learning algorithm. Models were trained using a subset of data available through MSHMP, evaluated for model accuracy and used to determine the relative importance of each predicting variable. Our results include a preliminarily ranking of variables based on their contribution to model predictions (Figure 1). Even after controlling for hog density and season of the year, we showed that the number of pigs received into neighboring farms was an important predictor of PED infection risk. These results indicate that the use of animal movement data as a neighborhood-level effect is innovative and will allow us to capture large-scale spatial dynamics based on long-distance movements to local spatial dynamics with neighborhoods of swine farms.
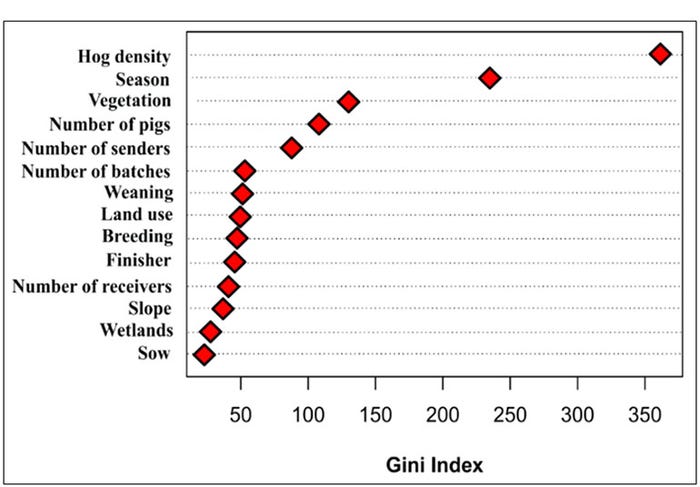
Figure 1: Relative importance in predicting risk according to the Gini index, a metric of information gain. “Number of pigs” represents the number of pigs received by a neighborhood during a three-month period.
References
1. Motta, P., Porphyre, T., Handel, I., Hamman, S. M., Ngwa, V. N., Tanya, V., Morgan, K., Christley, R. & Bronsvoort, B. M. D. (2017). Implications of the cattle trade network in Cameroon for regional disease prevention and control. Scientific Reports 7, doi:10.1038/srep43932
2. Lee, K., Polson, D., Lowe, E., Main, R., Holtkamp, D. & Martinez-Lopez, B. (2017). Unraveling the contact patterns and network structure of pig shipments in the United States and its association with porcine reproductive and respiratory syndrome virus (PRRSV) outbreaks. Preventive Veterinary Medicine 138, 113-123. doi.org/10.1016/j.prevetmed.2017.02.001
You May Also Like



