Genetics and early predictors of disease resilience
Selecting pigs for disease resilience can help reduce incidence, impact of infectious diseases.
March 2, 2021

Infectious disease represents one of the largest cost components to the swine industry, incurring veterinary costs, loss of pigs, reduced performance, and animal welfare. Strategies to reduce the incidence and impact of infectious disease include biosecurity, vaccination and routine use of medications. However, biosecurity protocols are not always effective, especially in hog-dense regions, and effective vaccines are not available for all pathogens. In addition, there is pressure from society to reduce the use of antibiotics in pork production.
An alternative strategy is to develop pigs that are genetically resistant to disease. However, complete resistance has been identified for only a few pathogens (e.g. E. coli F18). Resistant pigs could also be artificially created by gene editing, which was successful in creating pigs that are resistant to the PRRS virus. However, such pigs are not yet approved for entry into the food chain. Additionally, creating pigs that are resistant to the multitude of pathogens that are present in the swine industry, with new ones continually emerging, is not possible in the foreseeable future. Given these limitations, pigs will continue to be exposed to, and infected by, a range of pathogens.
It is, however, well known that some pigs fare better when exposed to disease than other pigs. This is called disease resilience; when exposed to disease, resilient pigs get less sick and/or they recover more quickly. And disease resilience is in part determined by genetics and can, thus, be selected for. The problem, however, is that breeding companies operate their selection programs in farms that are required to be free of major diseases, which makes it is impossible to directly identify and select more resilient pigs.
To better understand the genetic basis of disease resilience and to develop tools to select for disease resilience, a group of researchers from Iowa State University, the University of Alberta, and the University of Saskatchewan teamed up with a consortium of seven breeding companies (PigGen Canada) to establish a natural disease challenge at the Center de Développement du Porc du Québec (CDPQ) in Canada. The program was made possible through funding from Genome Canada and USDA.
The general design of the natural disease challenge is illustrated in Figure 1 and mimics a wean-to-finish farm with high disease pressure to maximize expression of genetic differences in resilience. The challenge environment was established by moving naturally infected animals (seeder pigs) into the facility, targeting various common viral and bacterial diseases. Subsequently, the challenge was maintained by introducing a new batch of 60 or 75 healthy nursery pigs every 3 weeks in a continuous flow system, allowing natural transmission from the previous batch to the new batch.
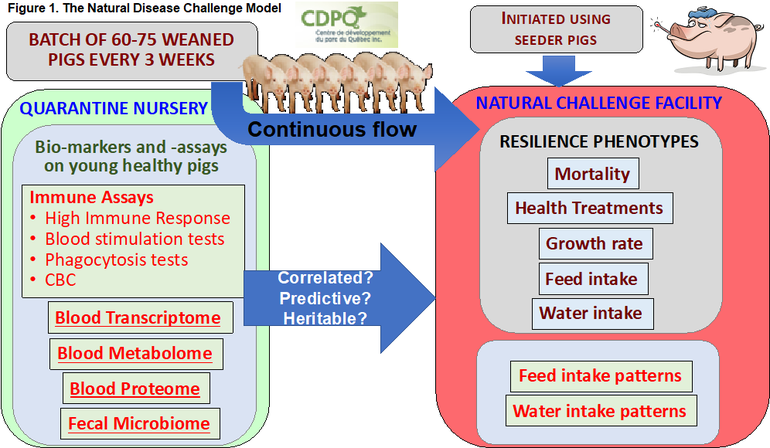
While in the challenge facility, detailed data are recorded on each pig, including veterinary treatments, mortality, growth rate, and individual feed and water intake, as well as carcass traits for pigs that survive to market. To date, we have analyzed data on over 3,200 pigs from 50 batches. Each batch of crossbred pigs was supplied by one of the seven PigGen companies from a multiplier in their system. It is important to note that the batch design does not allow us to compare the performance of pigs from different companies, and that is also not the purpose of this project. Instead, the goal is to identify differences in disease resilience between pigs from the same company.
Substantial animal-to-animal differences were observed for all traits collected in the challenge facility, and most traits were found to be heritable. This demonstrates that a pig’s resilience to this highly dynamic and relatively uncontrolled natural disease challenge is heritable and can be selected for. The key here is that what is measured is not a pig’s resilience to a specific disease but rather its resilience to the multiple pathogens that are present in this facility, similar to what would occur under a severe disease pressure in the field.
Another unique aspect of this study is that feed and water intake is recorded on every pig, using electronic equipment fitted with RFID readers. Figure 2 shows the day-to-day feed intake pattern of an example pig. Two big drops in feed intake can be seen that are very likely due to disease, as they coincided with medical treatments that the pig received. Based on these daily feed intake data, two indirect measures of disease resilience were developed: the proportion of “off-feed days” and “day-to-day variation in feed intake.” Off-feed days are indicated by red dots in Figure 2, based on the pig eating much less than expected (below the red line).
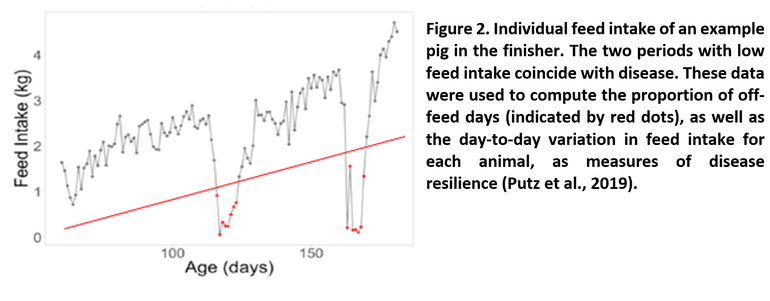
Genetic analyses found these and related feeding and drinking behavior traits to be moderately heritable and genetically correlated with disease resilience. This means that selecting pigs with a lower day-to-day variation in feed intake or a lower proportion of off-feed days is expected to improve disease resilience.
This, however, still leaves the question of how this could be applied in a high-health nucleus breeding program. Although extensive individual feed intake data is recorded in these programs, which could be used to measure day-to-day variation in feed intake or the proportion of off-feed days, these pigs are not exposed to major pathogens. Yet, other minor disease episodes do occur, and there are also other stressors such as heat stress and stress from interactions with other pigs and handling. These events may also be reflected in feed intake patterns.
To test whether day-to-day variation in feed intake without major disease indeed captures disease resilience, geneticists at Topigs Norsvin, one of the industry collaborators through PigGen Canada, recently computed day-to-day variation in feed intake based on their nucleus data (Harlizius et al. 2020). They found that progeny of boars that had lower estimated breeding values for day-to-day variation in feed intake indeed had lower mortality after they were experimentally infected with PRRS virus, followed by a natural challenge with numerous other pathogens. This suggests that breeding companies can use day-to-day variation in feed intake measured on their nucleus populations to select for increased resilience in the field.
Another solution to the problem of selecting for disease resilience in biosecure nucleus populations is to identify so-called “bio-markers” or “bio-assays” that can be measured on healthy pigs (preferably at a young age). For such bio-markers or bio-assays to be useful to select for disease resilience, they must be heritable and also genetically correlated to disease resilience. A blood sample can be a useful source to measure such bio-markers or bio-assays, as is a fecal sample.
To investigate this in the natural disease challenge project, blood and fecal samples were collected on all pigs before they entered the challenge (see the left part of Figure 1). The blood samples were evaluated for multiple assays to assess effectiveness of the pig’s immune system. Also, several high-throughput -omics technologies were applied to these samples, including transcriptomics to measure expression of over 15,000 genes, proteomics to measure over 400 proteins, and metabolomics to measure over 40 metabolites, as well as microbiomics to evaluate the fecal sample. Several of these bio-markers and -assays were found to have moderate heritability and to be genetically correlated with subsequent disease resilience of these same pigs in the natural disease challenge. This makes them good candidates to select for disease resilience among young healthy pigs, i.e. in nucleus breeding programs. There is also potential to use these bio-markers and -assays to predict the risk of disease of nursery pigs for management purposes or for “individualized” medicine.
Take home points:
If pigs can’t be made completely resistant to diseases, then at least make them more disease resilient, which is the ability of a pig to maintain productivity in the face of disease.
Genetic selection can play an important role in developing pigs that are more resilient to multiple diseases and other stressors.
Genetic selection for disease resilience, however, requires traits that can be measured on healthy pigs and that predict resilience of their offspring. Examples of such traits that were identified in this project are:
Day-to-day feed intake patterns, even recorded on healthy pigs
Bio-markers and bio-assays obtained from blood or fecal samples from young healthy pigs
References
Cheng, J., Putz, A.M., Harding, J.C., Dyck, M.K., Fortin, F., Plastow, G.S., Canada, P. and Dekkers, J.C., 2020. Genetic analysis of disease resilience in wean-to-finish pigs from a natural disease challenge model. Journal of Animal Science, 98(8), p.skaa244. doi.org/10.1093/jas/skaa244
Harlizius, B., Mathur, P. and Knol, E.F., 2020. Breeding for resilience: new opportunities in a modern pig breeding program. Journal of Animal Science, 98(Supplement_1), pp.S150-S154.
Putz, A.M., Harding, J.C., Dyck, M.K., Fortin, F., Plastow, G.S., Dekkers, J.C. and Canada, P., 2018. Novel resilience phenotypes using feed intake data from a natural disease challenge model in wean-to-finish pigs. Frontiers in genetics, 9. doi.org/10.3389/fgene.2018.00660 .
The natural challenge project was approved by the Animal Protection Committee of the Centre de Recherche en Sciences Animales de Deschambault (15PO283) and by the Animal Care and Use Committee at the University of Alberta (AUP00002227). The project was funded by Genome Canada, Genome Alberta, Genome Prairie, Alberta Pork, PigGen Canada, and USDA-NIFA grant # 2017-67007-26144. Members of PigGen Canada are acknowledged for providing the pigs and input into the project, including: Canadian Centre for Swine Improvement, Fast Genetics, Genesus, Hypor, ALPHAGENE, Topigs Norsvin, DNA Genetics, the Canadian Swine Breeders Association, and Alliance Genetics Canada.
Sources: Jack Dekkers, Jian Cheng, Fred Fortin, Mike Dyck, John Harding, PigGen Canada, Graham Plastow, Iowa State University, Centre de développement du porc du Québec inc., University of Alberta, University of Saskatchewan, who are solely responsible for the information provided, and wholly own the information. Informa Business Media and all its subsidiaries are not responsible for any of the content contained in this information asset.
You May Also Like



