Length of transmission and number of cases during spring raises concern about next season of PRRS.
August 10, 2021

During the fall of 2020, the simultaneous occurrence of farm-level PRRS outbreaks caused by unusually similar viruses at the orf5 level was reported by swine producers in the Midwestern U.S. Three main factors caused these farm-level outbreaks to quickly attract attention from the industry: 1) PRRSV with high orf5 nucleotide identities (>99%) were being detected in multiple apparently unrelated sites from different production systems; 2) this variant was affecting mainly growing pig sites and seemed different from variants historically detected in those production systems; and 3) reports from the field suggested that affected sites were experiencing extremely high production losses.
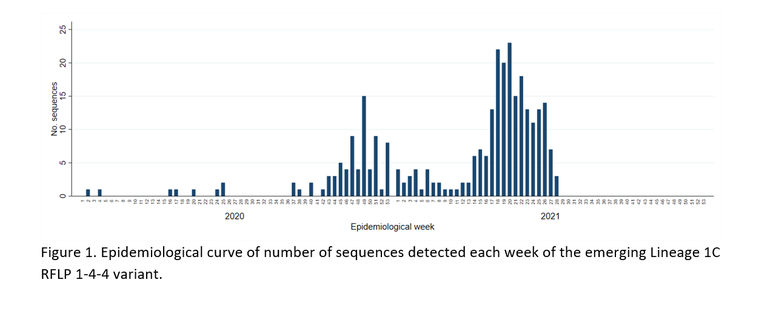
The Morrison Swine Health Monitoring Project (MSHMP) monitors disease occurrence in approximately 50% of the U.S. swine breeding herd. PRRSV orf5 sequences generated by the participating systems as a result of their routine outbreak investigations in breeding, gilt developing units, nursery, growing and finishing pig herds are captured. As of July 2021, a total of 301 sequences of this newly emergent L1C 1-4-4 variant from 14 MSHMP production systems was detected in 243 sites (163 growing pig farms, 53 breeding herds, 31 reported as other types, and 7 with no information on site type). The epidemiological curve of number of sequences detected per week is shown in Figure 1. We found that the transmission of this variant seemed to have occurred in two waves. The first one occurred between October and December 2020, when these cases first started being reported. This was followed by a decrease in the weekly cases in the next few months. However, by April 2021 a second wave of transmission was noted, peaking in May 2021. The number of sequences that comprised this second wave of transmission far surpasses the number of sequences observed during the fall-winter 2020 wave. Coordinates or state information were not available for 15% of cases, but this variant has been detected in Minn., Iowa, Ill., S.D. and Wis.
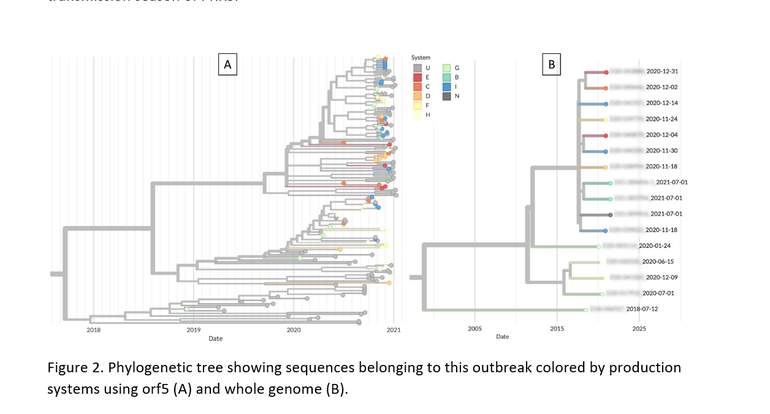
In order to evaluate if this genetic cluster represented the emergence of a new variant within its sub-lineage (1C), phylogenetic trees with orf5 (Figure 2A) and whole genome sequences from a subset of 17 cases (Figure 2B) wer constructed comparing these cases to other Lineage 1C sequences from MSHMP, the University of Minnesota Veterinary Diagnostic Laboratory and publicly available (GenBank). Both the orf5 and whole genome trees suggest that this is likely the emergence of a new PRRSV variant within Lineage 1C since no recent ancestor since approximately 2018 was found. Additionally, very similar sub-clade structures were found for this genetic cluster in both the orf5 and whole genome trees. This suggests that, at least for this newly emerging variant that is different from historical Lineage 1C PRRS sequences, interpretation of orf5 and whole genome remains the same in terms of all case sequences comprising their own separate clade and its sub-structures being preserved.
It is important to highlight, however, that this analysis is based on sequence data obtained from MSHMP participating systems and that non-MSHMP production systems may also be experiencing cases. Thus, although representative of U.S. swine population, the number of affected sites is likely higher than what was presented here. While the Midwest U.S. seems to have been the only heavily affected region by this PRRSV variant at the time of writing, the fact that transmission carried on throughout the spring and the number of cases surpassed the observed cases during winter raises concern about the next high transmission season of PRRS.
Source: Mariana Kikuti, Igor A. D. Paploski, Nakarin Pamornchainavakul, Catalina Picasso-Risso, Albert Rovira, Kimberly VanderWaal, Cesar A. Corzo, University of Minnesota, who are solely responsible for the information provided, and wholly own the information. Informa Business Media and all its subsidiaries are not responsible for any of the content contained in this information asset.
You May Also Like



