Hog Reproduction
thumbnail
Livestock Management
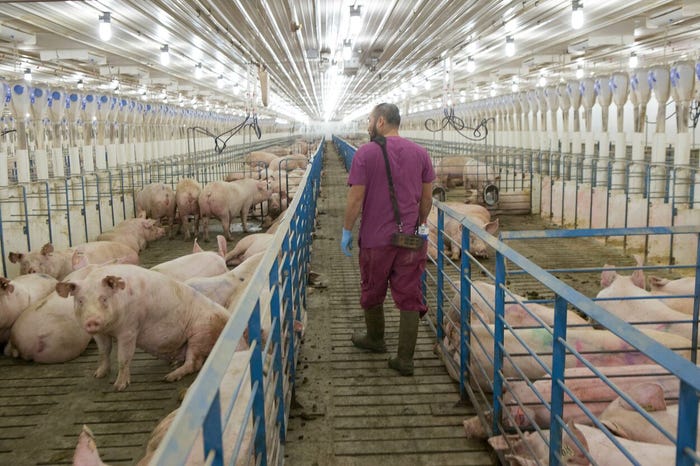
Maximizing pork production: Importance of a quality breed targetMaximizing pork production: Importance of a quality breed target
Sow farms that have the best opportunity for consistent farrowing rates, productivity, will drastically minimize any repeat services in their farms.
Subscribe to Our Newsletters
National Hog Farmer is the source for hog production, management and market news





















.jpg?width=300&auto=webp&quality=80&disable=upscale)






.jpg?width=300&auto=webp&quality=80&disable=upscale)