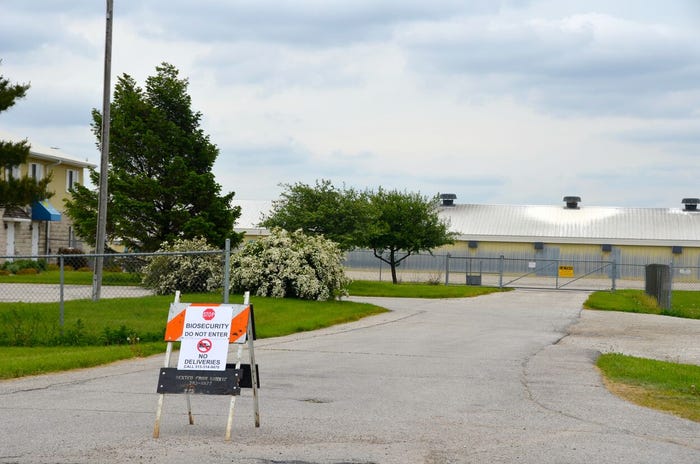
Biosecurity
thumbnail
Farming Business Management
USDA joins FarmRaise to offer online livestock disaster decision toolsUSDA joins FarmRaise to offer online livestock disaster decision tools
LIP also covers losses due to eligible diseases and attacks by animals reintroduced into the wild.
Subscribe to Our Newsletters
National Hog Farmer is the source for hog production, management and market news


.jpg?width=700&auto=webp&quality=80&disable=upscale)

























.jpg?width=300&auto=webp&quality=80&disable=upscale)

.jpg?width=300&auto=webp&quality=80&disable=upscale)



