North Carolina and Nebraska appear to have distinct clusters of PRRSV 1-7-4 strains.
February 5, 2019

By Giovani Trevisan, Aditi Sharma, Phillip Gauger, Chris Rademacher and Daniel Linhares, Iowa State University
The Swine Disease Reporting System database demonstrates since 2015 the porcine reproductive and respiratory syndrome virus restriction fragment length polymorphism type 1-7-4 has been the most common PRRSV wild type detected on cases submitted for open reading frame 5 sequencing at the Iowa State University -Veterinary Diagnostic Laboratory. The purpose of this analysis was to describe the nucleotide diversity of ORF-5 from PRRSV classified within the 1-7-4 RFLP.
Discriminant analysis of principal components [Fig. 1] was conducted for 769 PRRSV 1-7-4 ORF-5 sequences collected from January to November of 2018 from 15 states in the U.S. States with more than 10 sequences in the period were used in the analysis excluding sequences without a state designation. Iowa had the highest number of ORF-5 sequences in the database representing 40.6% of the total.
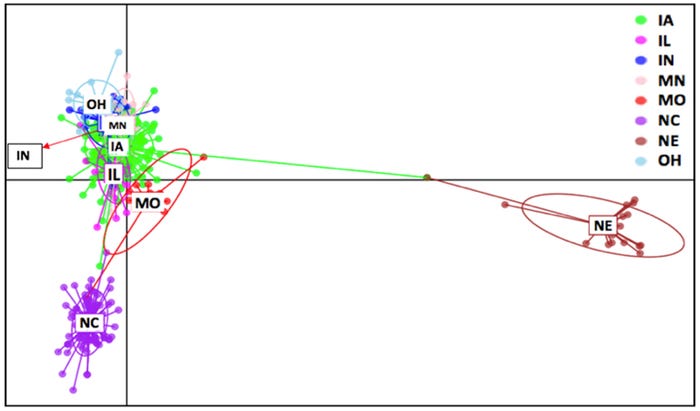
Figure 1: DAPC plot of ORF5 sequences for PRRSV RFLP type 1-7-4 for calendar year 2018 present at the SDRS database according to the state of origin. Each different color represents a different state. Each inertia ellipses shows a different cluster, while each dot represents individual sequences.
The DAPC plot [Fig. 1] demonstrates the genetic relatedness between Iowa, Minnesota, Indiana, Illinois and Ohio. Missouri had similar genetic diversity compared to the five previous states. In contrast, sequences from North Carolina and Nebraska were more genetically distant from all other states forming two distinct clusters. Distinct phylogenetic clusters in North Carolina and Nebraska could be due to the relative geographical and genetic isolation of the virus.
To further understand the genetic diversity among PRRSV sequences, pairwise nucleotide distances within and between states was calculated [Fig 2]. There is a considerable range of PRRSV 1-7-4 genetic diversity within and between states. The overall pairwise distances were ~ 2.8%. The most genetically distant PRRSV sequences differed by more than 8%. Interestingly, Nebraska and Missouri had the lowest percent genetic diversity.
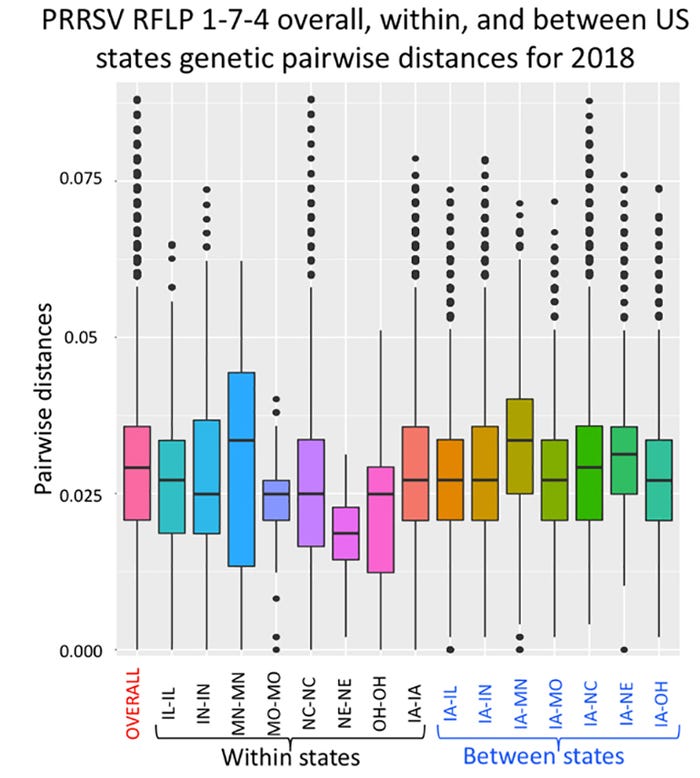
Figure 2: Box plot of pairwise ORF5 nucleotide distances for 769 PRRSV, within states, and between Iowa and other states. Each box-plot represents a different comparison. Box boundaries represent the second and third quartiles and the division inside the box the median. Lower and upper whiskers represent the first and fourth quartile. Open circles outside upper and lower whiskers represent outliers.
The RFLP is a historic method used to describe genetic relationships between PRRSV ORF5 sequences and are commonly reported by VDL’s. This analysis using the SDRS database for PRRSV 1-7-4 demonstrates substantial genetic diversity within this RFLP type that can be above 8% on a nucleotide basis. PRRSV from different states formed separate clusters with the majority demonstrating genetic relatedness, which could reflect the movement of pigs and corresponding PRRSV to different geographic regions in the U.S. Conducting a phylogenetic analysis of PRRSV detected within production systems will help monitor genetic diversity and better understand if the same or different/new PRRSV is circulating in the population.
Conclusions:
Within the PRRSV 1-7-4 strain there is great diversity. There were some individual sequences that showed pairwise difference close to 8%.
There is regional difference on genetic diversity of PRRSV 1-7-4.
North Carolina and Nebraska appear to have distinct clusters of PRRSV 1-7-4 strains. One hypothesis would be that these states do not import many pigs in comparison to the others.
Source: Iowa State University, who is solely responsible for the information provided, and wholly owns the information. Informa Business Media and all its subsidiaries are not responsible for any of the content contained in this information asset.
You May Also Like


