December 21, 2015

Porcine reproductive and respiratory syndrome virus costs U.S. hog producers substantially every year. According to a pork checkoff-Iowa State University study from 2011, the U.S. swine industry loses $664 million per year at the hands of PRRSV, with 45% of that loss attributable to the breeding herd.
Vaccines have proven beneficial, but do not provide 100% protection for heterologous strains, leaving producers to stick to biosecurity, herd elimination and regional vaccination as the preferred methods of control.
Infection with PRRSV causes reproductive failure in pregnant females. While in early gestation the virus can cause embryonic death, the clinical manifestation of PRRSV mainly occurs in late gestation and is characterized by abortions, early farrowings, fetal death and the birth of weak, congenitally infected piglets, resulting in elevated preweaning mortality.
Although transplacental PRRSV infection mainly occurs in late gestation, the exact mechanisms by which PRRSV transmits from the dam to her fetuses have yet to be determined. It has been suggested that fetal death may not be a direct result of PRRSV infection of fetal tissues since severe microscopic lesions are not observed in infected fetuses. It has been shown that the number of sialoadhesin-positive (CD169) and CD163-positive macrophages in endometrium and placenta, and virus replication in fetal implantation sites, which causes apoptosis of infected and surrounding cells, play a role in fetal death. Thus, it is postulated that fetal demise is mainly associated with events occurring at the maternal-fetal interface that lead to the separation of the fetal placenta from the uterus.
Advances in genomic technology and computing power have unleashed a variety of new tools that can be used to control diseases in human and animal populations. Genomic research has recently found novel genes associated with host responses to PRRSV and PCV2 in nursery pigs, and antibody responses in gestating sows.
Novel findings from the Pregnant Gilt Model will contribute to the development of genetic lines that are more resilient to reproductive PRRSV, helping to mitigate losses in the event of a reproductive outbreak.
The PGM involves the experimental infection at the University of Saskatchewan of more than 150 pregnant gilts with a highly virulent strain of PRRSV. The objectives were to improve understanding of the mechanisms resulting in fetal infection and death, and to investigate the genomic basis of resistance to reproductive PRRSV.
Undertaken in collaboration with a number of domestic and international research partners (Figure 1), the PGM included a pilot experiment (15 gilts) to develop laboratory methods and evaluate the relative virulence of three North American type 2 PRRSV strains. This was followed by the main experiment using purebred pregnant Landrace gilts that were selected on the basis of their average litter birth weight — about 50% were from low birth rate litters, and 50% were from high birth weight litters.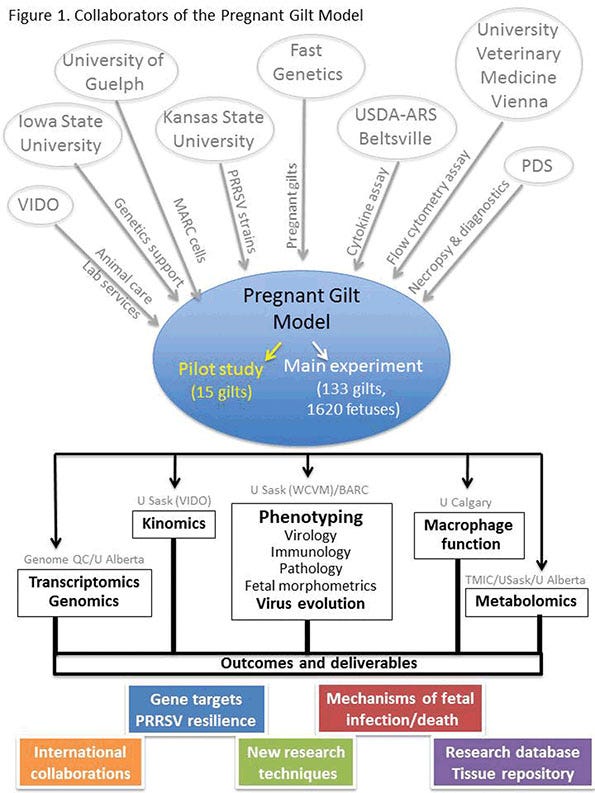
At gestation day 85 (±1), 114 gilts were experimentally infected with type 2 PRRSV. Nineteen control gilts were similarly mock-infected, and all gilts and fetuses were humanely euthanized at 21 days post inoculation (gestation day 106±1). Laboratory analyses of the estimated 50,000 samples collected enabled thorough evaluation of fetal and maternal host responses to PRRSV.
Clinical responses
Thirty-two percent of the inoculated gilts showed reduced daily feed intake, and 35 of the 111 inoculated gilts experienced fever versus none of the control gilts. One gilt died at 11 dpi and two gilts aborted at 17 and 20 dpi, respectively.
Excluding mummies, the average litter sizes were not statistically different between the groups, but the fetal mortality rate was significantly higher in the inoculated gilts — 41.0±22.8% compared to 1.4±3.4% in the non-inoculated gilts. Body weight of the viable fetuses from inoculated gilts was 17% lower than fetuses from non-inoculated dams. This study provided no substantive evidence that the severity of PRRS is influenced by dam birth weight; however, fetuses with intra-uterine growth retardation had lower PRRS viral loads in both fetal thymus and the maternal-fetal interface, suggesting that larger fetuses are at a higher risk of PRRSV infection and death.
Researchers also studied the viral and immunologic responses, pathologic responses, phenotypic predictors of reproductive PRRS severity and whole-blood transcriptomic responses.
RNA obtained from whole blood collected on 0, 2 and 6 dpi from a subset of gilts representing “resistant” (RES; n=8) and “susceptible” (SUS; n=8) gilts were used in a genome-wide analysis of the transcriptomic response to PRRSV infection. The RES group had a mean fetal mortality rate of 5% (range of 0 to 13%) compared to 76% (range of 67% to 94%) for the SUS group.
An RNA-sequencing approach with an Illumina sequencing platform was used to investigate differentially expressed genes between groups. DEGs were identified at all time points that were predictive of susceptibility to reproductive pathology. Fifteen genes were upregulated in RES gilts at all three time points, of which seven encode proteins with known functions in platelet cells. These cells function in repairing vascular injury and regulating the inflammatory and innate immune responses.
The RES gilts exhibited higher levels of interferon signaling at day 0 than SUS gilts, which could indicate that they were primed to respond faster to a PRRSV infection. This finding may offer the exciting prospect of identifying blood biomarkers that could be used to develop a screening test to predict the extent of fetal pathology in pregnant dams prior to PRRSV infection. By 2 dpi, the expression of pro-inflammatory genes in the RES group was lower than in SUS groups, while expression of T cell-related transcripts was higher. At day 6, interferon-gamma signaling was higher in RES than SUS pigs.
Overall, the expression data suggests that progression of the immune response to PRRSV is faster in RES than SUS dams, and that this may contribute to their lower levels of fetal pathology.
Novel findings
Novel findings include:
■ Viral load at the maternal-fetal interface and in fetal thymus at 21 dpi, but not in dams’ sera, was a strong predictor of the probability of fetal death.
■ Fetal infection and death clustered within the uterus, indicating the status of adjacent fetuses and inter-fetal transmission of PRRSV are important factors.
■ Gilts categorized as PRRS-resistant, based on having low fetal mortality and fetal viral load following challenge, had a faster immune response to PRRSV challenge than did gilts categorized as more susceptible.
■ The WUR10000125 SNP on chromosome 4, associated with lower PRRS viral load and higher average daily gain in experimentally infected nursery pigs (Boddicker, 2012), was not associated with reproductive outcome.
■ A genome-wide association study found 21 candidate genomic regions associated with fetal death, fetal viability or viral load in the thymus or the uterus.
The PGM was the largest study of reproductive PRRS ever undertaken, which enabled the use of a broad array of “omic” technologies (genomics, transcriptomics, proteomics, kinomics and metabolomics) to investigate phenotypic and genotypic predictors of the severity of reproductive PRRS. In-depth analyses led to the discovery of genomic and phenotypic mechanisms that may lead to novel control strategies for reproductive PRRS.
Building on these results and using a refined animal model and earlier termination, a new experiment, the PGM2, will validate the genomic markers associated with viral load and fetal autolysis, and investigate specific mechanisms underlying transplacental transmission and host responses occurring in the most susceptible fetuses.
To reiterate, PRRS is one of the most costly diseases affecting the global swine industry, and vaccines are not 100% effective. Alternative methods of control and prevention, such as identifying more resilient pigs, are warranted.
Contact John Harding for more information at [email protected]. Funding for the project was provided by grants from Genome Canada and Genome Prairie (Saskatchewan Ministry of Agriculture and Food), with administrative support from Genome Alberta.
You May Also Like



