May 27, 2016

Respiratory disease is a major health concern in swine production in the United States. Several viruses play a role in the porcine respiratory disease complex including porcine reproductive and respiratory syndrome virus, influenza A virus and porcine circovirus type 2, although infections often involve more than one of these respiratory pathogens.
The Iowa State University Veterinary Diagnostic Laboratory received submissions of respiratory disease from pigs demonstrating lethargy, coughing and sneezing with microscopic lesions consistent with bronchointerstitial pneumonia. Lung tissue also exhibited bronchiolitis with bronchiolar epithelial necrosis, a common lesion caused by infection with influenza A virus.
Lung tissue, nasal swabs and oral fluids were polymerase chain reaction-negative for pathogens commonly associated with respiratory disease in pigs prompting further investigation. Additional diagnostics using broad genus-level PCR assays detected a sequence with genetic similarity to a porcine parainfluenza virus type 1 reported in Hong Kong suggesting an association between this Paramyxovirus and respiratory disease in swine.
Paramyxoviruses are pathogens known to infect humans and livestock species including swine. The proposed porcine parainfluenza virus type 1 belongs to the genus Respirovirus in the subfamily Paramyxovirinae within the family Paramyxoviridae. In 2013, PPIV-1 was reported in nasopharyngeal samples collected from slaughterhouse pigs in Hong Kong, China. Yoon reported finding cases in the United States in March 2013 using a broad genus-level PCR assay.
Although diagnostic laboratories have detected PPIV-1 in diagnostic samples, information regarding the epidemiology, transmission, lesions and clinical disease associated with PPIV-1 is lacking. However, the utility of metagenomics and next generation sequencing has increased our ability to detect unknown viruses and accumulate sequence information to improve diagnostic capabilities.
PPIV-1 has been detected in oral fluids, nasal swabs and lung tissue by PCR. The ISU VDL recently performed PPIV-1 PCR testing that was done on samples collected in February-to-April of 2016 and found that 38% of samples tested were positive for PPIV-1 (Figure 1). 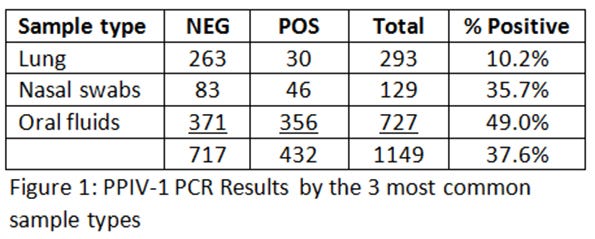
The vast majority of these samples are oral fluids and nasal swabs. Most of the samples did not have the ages included, but a visual assessment of the data would indicate that most of the positive samples were found anywhere from 2 to 15 weeks of age. This would seem to corroborate published and anecdotal reports of finding PPIV-1 late in lactation (nagging cough in the piglets in the last week of lactation) or early in the post-weaning phase where influenza A virus has been ruled out, but had lesions that would be consistent with influenza.
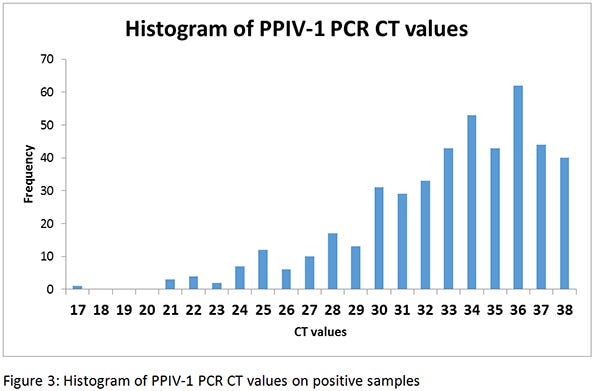
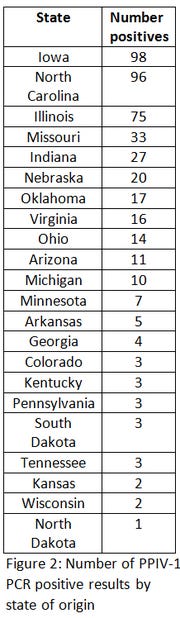 Figure 2 shows the distribution of states where the PPIV-1 PCR positive samples were found, and it clearly shows that the virus appears to be widespread throughout the swine industry, since it was located in 22 states in this small dataset. Although the virus appears to be fairly prevalent, an analysis of the cycle threshold values do not seem to indicate the virus is present in high quantities (high CT values), although this would certainly be influenced by the sample type (Figure 3).
Figure 2 shows the distribution of states where the PPIV-1 PCR positive samples were found, and it clearly shows that the virus appears to be widespread throughout the swine industry, since it was located in 22 states in this small dataset. Although the virus appears to be fairly prevalent, an analysis of the cycle threshold values do not seem to indicate the virus is present in high quantities (high CT values), although this would certainly be influenced by the sample type (Figure 3).
A recent report from Kansas State University used in situ hybridization to demonstrate PPIV-1 genetic material in nasal turbinate and to a lesser extent in trachea suggesting the virus replicates in the upper respiratory tract. Unfortunately, clinical disease and lung lesions associated with PPIV-1 have not been thoroughly reported due to the lack of controlled experiments. Attempts to isolate the virus have been unsuccessful to date precluding the ability to evaluate PPIV-1 through experimental infection. However, detection of the virus by PCR in diagnostic submissions suggest the virus is widespread in the swine population.
You May Also Like



