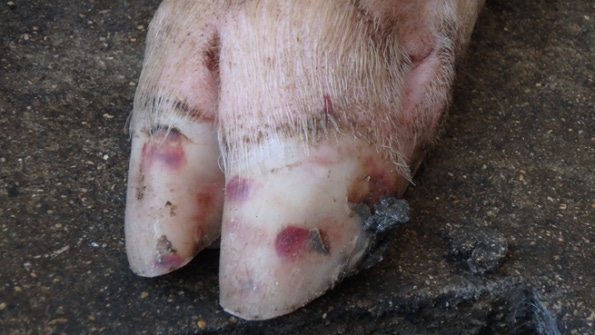
The primary concern has been Senecavirus A-induced vesicles on pigs’ snout and feet look exactly like the lesions produced by our most important foreign animal diseases.
December 1, 2016
Senecavirus A, an RNA virus, was first described in the United States in the late-1980s, but was not associated with clinical disease until 2007 (“idiopathic vesicular disease”). Highly sporadic and transient, idiopathic vesicular disease was also occasionally reported in pigs in Australia, New Zealand and Canada (16, 8).
In 2014, SVA was detected in association with outbreaks of vesicular disease in combination with neonatal mortality in the United States, Brazil and China (5, 10, 11, 12). Affected breeding herds reported neonatal morbidity and mortality ranging from 30 to 70%, primarily in piglets 7 days of age or younger. In these cases, neonates did not present with specific clinical signs or lesions, but SVA could be detected in a variety of tissues collected from piglets.
The lesions associated with SVA are characterized by vesicles and epidermal erosions that progress to ulcers of the coronary band, oral cavity and nasal planum. Affected animals present transient fever and lameness. Lesions and clinical signs have been reproduced in finisher pigs inoculated with SVA under experimental conditions (3, 7). In contrast, the neonatal mortality associated with SVA in some sow farms has not been replicated experimentally.
Although formal studies have not been done to estimate the costs of an SVA outbreak, recent cases we investigated suggested that SVA can cause substantial losses in neonatal, grow-finish and gestating pigs. Most commonly, the primary concern has been SVA-induced vesicles on the snout and feet that look exactly like the lesions produced by our most important foreign animal diseases: foot-and-mouth disease virus, vesicular stomatitis virus, swine vesicular disease virus and vesicular exanthema of swine virus. FAD investigations and diagnostic testing must be done whenever vesicles are observed to guard against the entry FMDV and other important FADs. In market-ready finishers, this has resulted in inconvenient shipping delays, but the consequences of an error would be catastrophic to the industry.
Similar to the pattern observed in 2015, 2016 saw an uptick in confirmed SVA cases beginning in mid-summer that has persisted through the fall (see Figure). If the 2015 pattern holds, we may see a decline in cases as cold weather sets in. The reasons for the apparent seasonality of clinical SVA have not yet been established.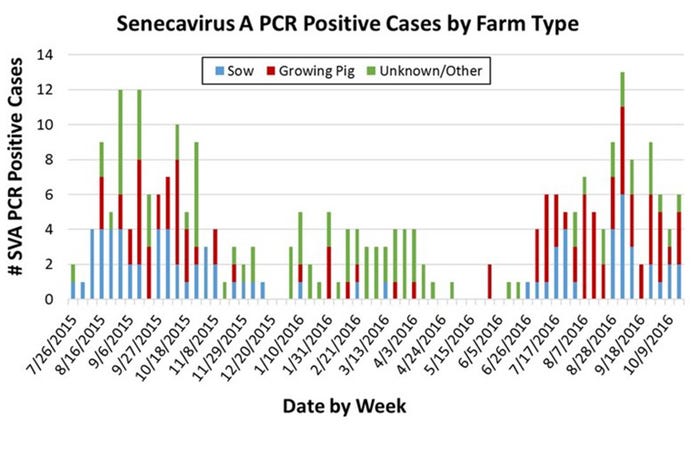
Phylogenetic analyses using complete genome sequences of contemporary SVA isolates and historical sequences suggest that SVA is diverse and continuously evolving — as would be expected of an RNA virus. Contemporary U.S. isolates share 98-99% nucleotide identity among themselves. By comparison, contemporary U.S. isolates share 91-93% nucleotide identity with the prototype U.S. SVA strain SVV001 and a Canadian isolate from 2007, 95-97% nucleotide identity with contemporary Brazilian isolates, and 94-96% nucleotide identity with a recent Chinese isolate (3, 4, 12). The implication of SVA viral diversity is not yet clear.
A diagnosis of SVA infection is based on clinical signs and diagnostic testing. Diagnostic assays currently available include virus isolation, detection of nucleic acid by PCR (targeting VP1 or 5’UTR), and direct detection in tissues by immunohistochemistry or in situ hybridization. SVA serum antibody assays include a recombinant VP1 ELISA, immunofluorescence, and virus neutralization.
The challenge in diagnosing SVA infection is the transient nature of the clinical signs and vesicular lesions. That is, their diagnostic value decreases rapidly over time. Joshi et al (2016) described a short viremia of three to seven days and viral shedding for up to 28 days. However, detection of SVA by PCR has been inconsistent in sows, suggesting variation in the pattern of shedding and/or the amount of virus sequestered in each type of specimen (2). Therefore, when faced with a situation suggestive of SVA, a variety of specimens (serum, tonsil, feces, oral fluid and vesicular fluid) should be collected for SVA PCR testing (9).
Antibody detection can be very helpful in establishing a SVA diagnosis. SVA encodes a polyprotein that contains four structural (VP1 to 4) and seven non-structural proteins. Among viruses related to SVA (picornaviruses), VP1 is the most immunogenic. Therefore, we developed an indirect SVA serum antibody ELISA using recombinant VP1 protein. In the field, this ELISA detected seroconversion against SVA in both sows with, and sows without, clinical signs of SVA at early stages of the outbreak. Likewise, maternal SVA antibodies were detected in serum collected from offspring (2).
The transience and variability of clinical signs, absence of vesicles (gross lesions) in many SVA-infected animals, and the inconsistency or variability of PCR results among specimens presents a challenge to diagnosticians. Reliance on a single diagnostic method will not achieve the necessary accuracy. Therefore, we recommend that the diagnosis of SVA be based on a combination of clinical observations, PCR testing of a variety of diagnostic specimens, and serum antibody testing.
References cited
1. Amass SF, Schneider JL, Miller CA, et al. 2004. Idiopathic vesicular disease in a swine herd in Indiana. J Swine Health Prod 12:192-196.
2. Gimenez-Lirola LG, Rademacher C, Linhares D, et al. 2016. Serological and molecular detection of Senecavirus A associated with an outbreak of swine idiopathic vesicular disease and neonatal mortality. J Clin Microbiol 54:2082-9.
3. Joshi LR, Mohr KA, Clement T, et al. 2016. Detection of the emerging picornavirus Senecavirus A in pigs, mice, and houseflies. 2016. J Clin Microbiol 54:1536-1545.
4. Laguardia-Nascimento M, Gasparini MR, Sales ÉB, et al. 2016. Molecular epidemiology of Senecavirus A associated with vesicular disease in pigs in Brazil. Vet J 216:207-209.
5. Leme RA, Zotti E, Alcântara BK, et al. 2015. Senecavirus A: An emerging vesicular infection in Brazilian pig herds. Transbound Emerg Dis 62:603-611.
6. Montgomery JF, Oliver RE, Poole WSH. 1987. A vesiculo-bullous disease in pigs resembling foot and mouth disease I. Field cases. New Zealand Vet J 35:21-26.
7. Montiel N, Buckley A, Guo B, et al. 2016. Vesicular Disease in 9-week-old pigs experimentally infected with Senecavirus A. Emerg Infect Dis 22:1246-1248.
8. Munday BL, Ryan FB. 1982. Vesicular lesions in swine - possible association with the feeding of marine products. Australian Vet J 59:193-193.
9. Piñeyro PE, Rademacher CJ, Derscheid RJ, et al. 2015. Senecavirus A vesicular disease outbreak in Midwest show and commercial pigs. Proceedings of the 58th of the American Association of Veterinary Laboratory Diagnosticians, Providence, RI.
10. Singh K, Corner S, Clark S, et al. 2012. Seneca Valley virus and vesicular lesions in a pig with idiopathic vesicular disease. J Vet Sci Technol 3:123.
11. Vannucci FA, Linhares DC, Barcellos DE, et al. 2015. Identification and complete genome of Seneca Valley virus in vesicular fluid and sera of pigs affected with idiopathic vesicular disease, Brazil. Transbound Emerg Dis 62:589-593.
12. Wu Q, Zhao X, Chen Y, et al. 2016. Complete genome sequence of Seneca Valley Virus CH-01-2015 Identified in China. Genome Announcements 4.
You May Also Like


