Understanding how the microbiome impacts health and disease of swine is an emerging area of study.
December 6, 2016

New research at the Kansas State University will change how the swine industry views the importance of gut health in a PRRS-endemic herd. Microbiome is a term used to describe the community of microorganisms that live on the skin and mucosal surfaces of animals. The gut microbiome houses the majority of these microorganisms and is critical for proper immune development and digestion of nutrients. Understanding how the microbiome impacts health and disease of swine is an emerging area of study.
Alternatives to antimicrobials are necessary for the management of infectious diseases in food producing animals. One potential alternative is the microbiome, used to describe the community of microorganisms that live within the digestive tract. Certain microbial populations have been associated with weight gain as well as response to infectious disease in humans and animals. For example, changes in the composition and diversity of the microbiome have been associated with pathogen load, inflammation, and muscle wasting in models of human infectious disease. How the microbiome is associated with systemic infections in swine is poorly understood.
In this research led by Megan C. Niederwerder, scientists investigate how the microbiome is associated with outcome in nursery pigs following co-infection with porcine reproductive and respiratory syndrome virus and porcine circovirus type 2. PRRS is the most costly disease of swine production worldwide, estimated to cost the U.S. swine industry approximately $664 million annual. After decades of research into disease control, PRRS continues to plague the industry and alternative approaches are necessary to reduce PRRS-associated losses.
Several causes of infectious pneumonia in humans have been found to have associations with the gut microbiome; however, very little information is available on how the gut microbiome may impact respiratory infections in pigs. They hypothesized that microbial colonization in the pig gut would influence response to porcine respiratory disease. The overall goal of this research was to identify characteristics of the pig microbiome that played a beneficial or detrimental role in clinical disease outcome following PRRSV/PCV2 co-infection.
This study provides new evidence on how microorganisms in the pig gut impact overall health and growth during virus infections. It was discovered that both microbial diversity and composition were associated with outcome following co-infection with PRRSV and PCV2.
Research method
An experimental population of 95 nursery pigs was co-infected with PRRS andPCV2 and followed for 70 days. The 4 week-old barrows were obtained from a single commercial source herd negative for PRRSV. The sow herd had no history of clinical disease associated with PCV2 or PRRSV. However, a relevant health history included a recent introduction of porcine epidemic diarrhea virus prior to farrowing. All piglets were confirmed to be negative for PRRS prior to being challenged. During the experiment all pigs were housed in environmentally controlled room at the Kansas State University Large Animal Research Center in 10 pens with nine or 10 pigs in each.
At the conclusion of the trial, 20 pigs were selected to represent the best and worst clinical outcomes in the co-infected population. Significant variation in weight gain and clinical disease occurred in the cohort of co-infected pigs. Best clinical outcome pigs had a complete lack of overt clinical signs and the highest average daily gain post-infection. Worst clinical outcome pigs had 10 or more days of moderate to severe clinical signs and the lowest ADG post-infection (refer to Figure 1). 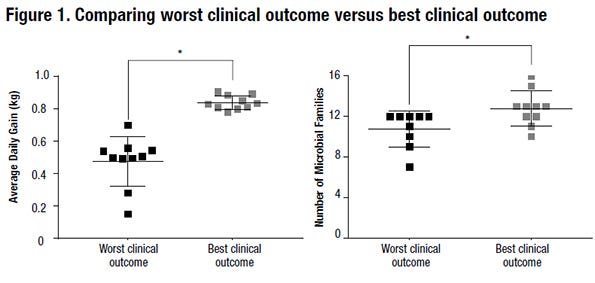
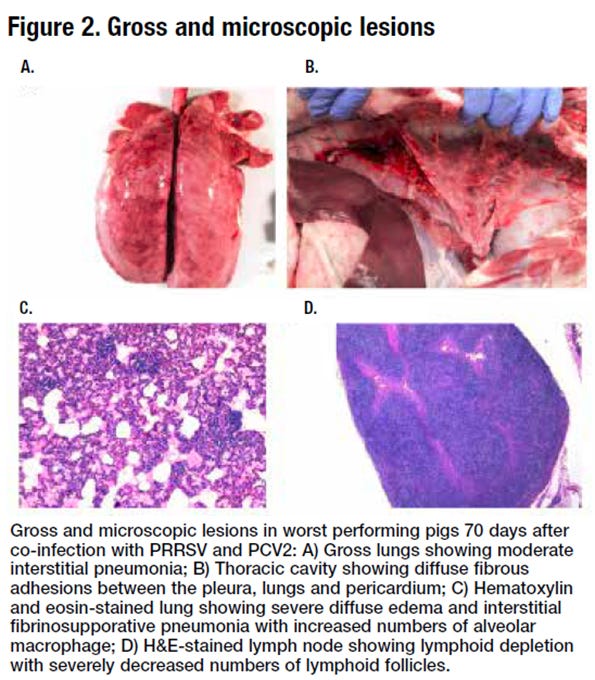
Best clinical outcome pigs had less interstitial pneumonia, fewer gross necropsy lesions, and lower levels of PRRSV and PCV2 in the blood. The fecal microbiomes of the two clinical outcome groups were evaluated at the conclusion of the trial to assess how microbial colonization of the gut may have played a role in determining outcome. The gross and microscopic lesions in worst performing pigs are shown in Figure 2.
Two significantly different microbiome characteristics were detected between the two outcomes. First, best clinical outcome pigs had increased microbial family diversity (refer to Figure 1). Second, best clinical outcome pigs had a higher prevalence of non-pathogenic Escherichia coli. This work provides the first evidence that the microbiome plays a role in outcome following co-infection with two common viral respiratory pathogens in swine. Moreover, this work suggests that the microbiome may serve as a valuable tool to improve the response to polymicrobial infections in growing pigs.
Improved outcome after co-infection was associated with reduced virus replication, increased weight gain, decreased clinical signs, and reduced interstitial pneumonia. With regards to the microbiome, improved outcome was associated with increased microbial diversity and increased prevalence of non-pathogenic Escherichia coli. These results suggest that the microbiome plays a role in systemic viral infections of swine and those certain microbial characteristics may be beneficial to outcome. Because these microorganisms are relatively easy to access and modify through the diet, the microbiome may be a novel tool for managing population diseases such as PRRS.
Although the science is in the early stages of learning how the microorganisms living inside the intestine impact health and disease throughout the whole pig, researchers know these microorganisms play a role in weight gain and immune response to pathogens. However, it was unknown if and how these microorganisms impact response to PRRS.
Niederwerder, M.C., Jaing, C.J., Thissen, J.B., Cino-Ozuna, A.G., McLoughlin, K.S., Rowland, R.R., 2016. Microbiome associations in pigs with the best and worst clinical outcomes following co-infection with porcine reproductive and respiratory syndrome virus (PRRSV) and porcine circovirus type 2 (PCV2). Veterinary microbiology 188, 1-11. For more information contact Megan Niederwerder at 785-532-4663 or [email protected].
This work was supported by the USDA NIFA Award #2013-68004-20362, the State of Kansas National Bio and Agro-Defense Facility Fund, and by the Lawrence Livermore National Laboratory Derived Research and Development effort (14ERD081).
You May Also Like



