Microbial communities residing in animals impact multiple aspects of the swine production system. Yet, it seems that producers still lack a complete picture of why funding microbiome research is critical to advance the pork industry.
March 15, 2018

By Andres Gomez (Animal Microbiomics), Pedro Urriola, Jerry Shurson (Swine Nutrition) and Lee Johnston (Swine Production), University of Minnesota Department of Animal Science
Microbes have been traditionally viewed through a lens of distrust, as pathogens affecting health. However, molecular and computational breakthroughs to study microbial diversity and function by sorting DNA sequences have presented a novel concept of an animal “flora” that acts as a friend as opposed to a foe.
Nowadays, we can sort and sequence the DNA of trillions of microbes in hundreds of subjects simultaneously. This tremendous ability has given us a snapshot of which microbes reside in animals and how they influence animal nutrition, performance, health, welfare and behavior. Knowledge of the diversity of the swine microbiome, the collection of bacteria (primarily), fungi, virus and protozoa colonizing the animal anatomy, can provide a novel, alternative perspective on the biological process that impacts some of the most important issues in swine production systems.
For instance, characterizing the specific microbes that increase or decrease in abundance upon pharmaceutical or dietary interventions is critical to determine precise dose-response relationships. Thus, if we identify patterns of change in the abundance of specific bacteria in response to particular feed additives in the gut, and correlate those microbiome changes with a trait of interest (e.g., feed-to-gain ratio), we can make more targeted and strategic use of feed additives, to potentially reduce feed costs while achieving desired improvements in pig health and productivity. Although, these kinds of studies have been performed previously, there has not been a systematic analysis to determine how specific nutrient fractions and quantities alter particular microbes and their ultimate impacts on productivity.
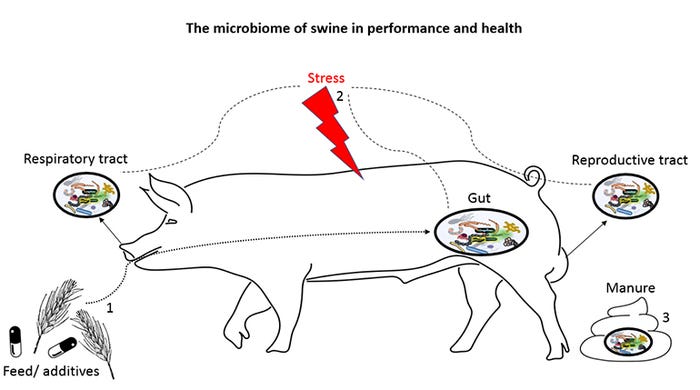
Events that affect swine performance and health through the microbiome: 1) nutrients and feed additives modulate gut microbiomes to impact health and performance, 2) microbiomes across the pig anatomy are accurate biomarkers of stress such as diseases, early weaning, and heat, and 3) microbiomes in manure can be modulated to mitigate harmful gases.
Likewise, defining “healthy” microbiomes within a population can be accomplished by making comparisons in microbiome diversity between good, average and poor performing pigs on a farm. This would allow producers to make early decisions on therapeutic or dietary interventions to enhance performance and health. Thus, regular “microbiome snapshots” along the most critical stages of pig growth (e.g., pre- and post-weaning), can be used to predict health and potential pathogen threats for disease by early identification of bacteria in slow-growing pigs or those that are at most risk of infection. Even though microbiome research in swine production systems has demonstrated an altered microbiome in poor performing or sick pigs, more studies are needed that better characterize the systematic changes in microbiome diversity over time (from birth to after weaning) to predict health and performance outcomes. Moreover, it is still unclear how the microbiome interacts with infectious agents (like viruses) to make pigs more susceptible to disease.
With the current regulations involving antibiotic use in the U.S. pork industry, alternatives to using growth-promoting antibiotics must be evaluated based on their impact on microbiome composition and function. This information must also be integrated with other systemic physiological responses to explain the mechanism of improving pig health and growth performance. For example, the microbiome in the gut or respiratory tract is a protective layer against infectious diseases. Thus, with microbiome research, we can determine how novel feed additives and management interventions work, by either enhancing the abundance of microbes that promote health and/or displacing those that cause disease. Similarly, because groups of microbes in a balanced microbiome compete with or kill potential pathogens or food safety hazards (e.g., Salmonella), microbiome research can help us identify novel approaches to improve food safety in the pork industry.
Microbiome research also extends to other processes in the pork production chain beyond the animal. For instance, specialized bacteria and fungi can degrade otherwise underutilized natural resources (e.g., dietary fiber in distillers dried grains with solubles) to maximize pig productivity, while decreasing the environmental footprint. Specialized microbial communities can also mitigate the production of dangerous gases (e.g. hydrogen sulfide) produced in manure pits. In this process beneficial bacteria overcome the competition from gas producers, leading to improved safety for pigs and workers in pork production facilities. Thus, we can improve many aspects of pork production systems by discovering and understanding the presence, function and abundance of various microbes with specialized roles in these processes. This kind of knowledge can only be generated by the molecular and computational tools that define microbiome science today.
In summary, the science for improving health, productivity, efficiency and safety of pork production systems can be greatly enhanced by conducting basic and applied microbiome research. The future of microbiome research will rely on integrating microbiome data with information on gastrointestinal physiology, metabolites and functional genomics in the whole animal. This approach will facilitate breakthroughs in elucidating the mechanisms of how various feed additives and nutrients interact with microbes and how microbes and the compounds they produce impact the animal. Similarly, determining the intrinsic (genetic) and environmental factors that define the core microbial communities across populations and breeds will help producers know the extent to which the microbiome can be modulated or selected to optimize the efficiency of pork production systems.
Gomez, Urriola and Shurson are members of the Integrated Animal Systems Biology Team at the University of Minnesota.
You May Also Like



