Collection of oral fluid samples at the abattoir is easier, more practical and cheaper than going farm-to-farm to collect samples for surveillance.
March 7, 2017

By Marcelo Almeida, Jeff Zimmerman and Daniel Linhares, Iowa State University College of Veterinary Medicine Veterinary Diagnostic and Production Animal Department
Eliminating porcine reproductive and respiratory syndrome virus from the U.S. swine industry would save up to $25 per finishing pig. However, achieving the goal of regional/national elimination programs requires implementation of a simple, reliable, cost-efficient surveillance system capable of providing producers the information they need to make informed disease control decisions: Should I vaccinate my finishing pigs? Where should I direct flows? Is my current control plan effective in controlling PRRSV in my downstream flow?
There is currently no active surveillance for PRRSV, chiefly due to the inconvenience and cost of conventional monitoring programs based on bleeding multiple pigs per site. Alternatively, a PRRS monitoring system based on collecting oral fluids at the abattoir, a point of concentration for finishing pigs, could be a practical and affordable method to track virus activity over time and geographical space.
To test this idea, we assessed the feasibility of using oral fluid samples collected from pigs at the abattoir to monitor 1) PRRSV exposure status by ELISA; 2) PRRSV circulation by real time quantitative molecular methods (qPCR). Results from abattoir samples were compared to those collected at the farm (within two days of shipping pig lots to slaughter).

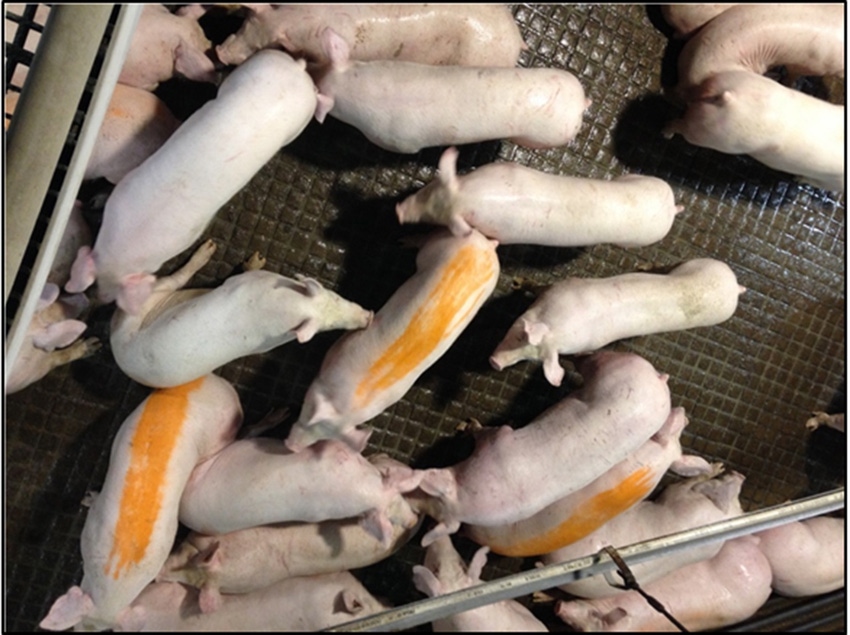
Sampling at farm (pens) Sampling at abattoir (lairage)
Methods
Serum (n=10) and oral fluid (n=10) samples were collected at finishing sites from 36 lots of pigs. Within 48 hours of collecting samples on the farm, the same pigs were followed to the abattoir where oral fluids samples (n=3) were collected while the pigs were in lairage. Samples were tested at the Iowa State University Veterinary Diagnostic Lab (Figure 1) for PRRSV RNA (polymerase chain reaction) and PRRSV antibody (ELISA). Thereafter, we compare the results from samples collected on the farm to samples collected at the abattoir.
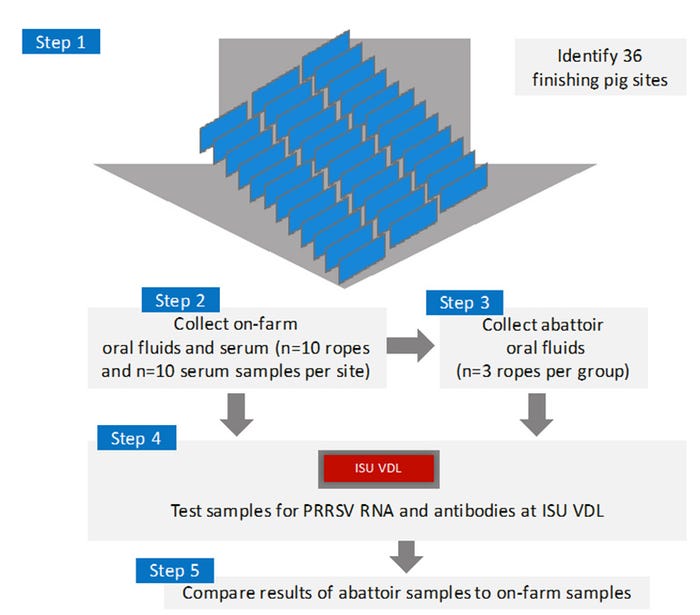
Figure 1: Experimental design: Assessment of PRRSV oral-fluid based abattoir surveillance.
Results and discussion
Oral fluids were successfully collected at the abattoir. Specifically, from 36 pig lots sampled at the farm (serum and oral fluids), oral fluid samples were collected from 32 groups of pigs (89%) at the abattoir. Anti-PRRSV antibodies were found in 93% of the serum samples and in all oral fluids (farm and abattoir). PRRSV RNA was detected in about 2% of sera and ~18% on oral fluids. Nine pig lots (28%) were detected as positive on farm, but were either suspect or negative at the abattoir. This discrepancy between group status may be related to the number of samples collected on farm (n=10 oral fluids) and at abattoir (three oral fluids), as it was observed that the higher the proportion of farm positive oral fluids, the higher it was the probability of detecting a group as positive at the abattoir (data not shown).
In summary, collection of samples at the abattoir is easier, more practical and cheaper than going from farm-to-farm to collect samples for surveillance. Furthermore, oral fluids, an aggregate sample, easy to collect, welfare friendly, and population based demonstrated to detect a higher number of animals as positive both in ELISA and PCR compared to serum samples. There is a growing number of oral fluids-based tests to detect antibodies or nucleic acids from endemic pathogens (including porcine epidemic diarrhea virus, Actinobacillus pleuropneumoniae, Influenza A virus) and foreign animal diseases (including foot and mouth disease, classical swine fever and African swine fever).
Moreover, PRRS control and elimination from swine populations require an active and efficient large-scale surveillance system. Validating oral fluids-based surveillance systems to detect PRRSV and other pathogens will make endemic disease elimination a possible task, which will significantly improve the global competitive advantage of the U.S. swine industry. Further studies are needed to characterize factors associated with farm-abattoir agreement of diagnostic results.
Acknowledgments
The implementation of this project was possible due to support from Drs. Donovan and Dion from The Hanor Co., Dr. Murray from New Fashion Pork and Dr. Wiseman from Triumph Foods LLC. We also thank Dr. Main from ISU-VDL, Dr. Torrison from the University of Minnesota and Dr. Christopher-Hennings from South Dakota State University. Likewise, we acknowledge Drs. Lopez and de-Souza-e-Silva from ISU Veterinary Diagnostic and Production Animal Medicine for the significant field support. Funding was obtained from the Swine Health Information Center.
You May Also Like



