One application would be to avoid mixing cohort group of piglets with low Ct values with piglets of different Ct values when commingling for nursery placement.
December 4, 2018

By Giovani Trevisan, Rebecca Robbins, Jose Angulo and Daniel Linhares, Iowa State University
Porcine reproductive and respiratory syndrome virus is the most economically significant pathogen in the U.S. swine industry1. The ability to monitor PRRSV status and virus circulation within a herd give a competitive edge to producers for proactively implementing management measures to reduce the losses caused by the infection.
One practical way to monitor PRRSV is by collecting the processing fluids which are generated from castration and tail docking, and testing by real-time reverse transcription-polymerase chain reaction for PRRSV RNA2. Personal communications with field veterinarians and farm personnel indicate that processing fluid is well accepted for routine sampling of the pre-wean piglet population. Traditional method of sampling piglets relied on American Association of Swine Veterinarians criteria that suggest the collection of 30 serum samples per month on due-to-wean piglets3. Processing fluid allows to sample larger numbers of piglets since every processed piglet could potentially contribute for the aggregated final sample. Data from the Swine Disease Reporting System show that during the year of 2018 processing fluids represented 9.4% of all cases tested for PRRS PCR by the four major swine veterinary diagnostic laboratories4, 5 indicating a rapid adoption of this specimen by the swine industry to monitor herds for PRRSV. A practical question is how PRRS PCR results on processing fluids relate to downstream performance parameters.
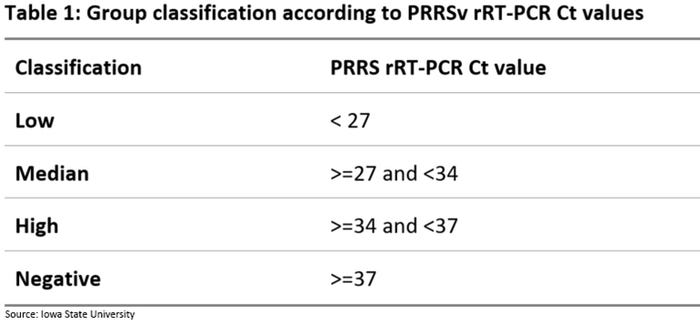
We conducted an observational study to investigate the relationship between processing fluid results for PRRSV tested by rRT-PCR and subsequent nursery mortality. Twelve sow farms routinely vaccinated with PRRS modified live virus vaccine were enrolled in this study. Collection of processing fluids from cohort groups took place during the three days of the week when the highest number of litters were processed. Samples were submitted to test for PRRSV RNA by rRT-PCR at the Iowa State University Veterinary Diagnostic Laboratory. Multiple processing fluid samples within a cohort group were pooled by 3:1 according to sow farm of origin and week of collection. The PCR Ct values were used to classify each piglet cohort group into four categories: Low, Median, High and Negative (Table 1).
Preliminary results include data from 126 cohorts. Piglet cohorts from low group had almost 3 percentage points higher mortality when compared to other groups. This difference was statistically significant having an ANOVA Tukey post-hoc adjusted p-value less than 0.05. No statistical difference was found between groups classified as median, high or negative (Table 2).
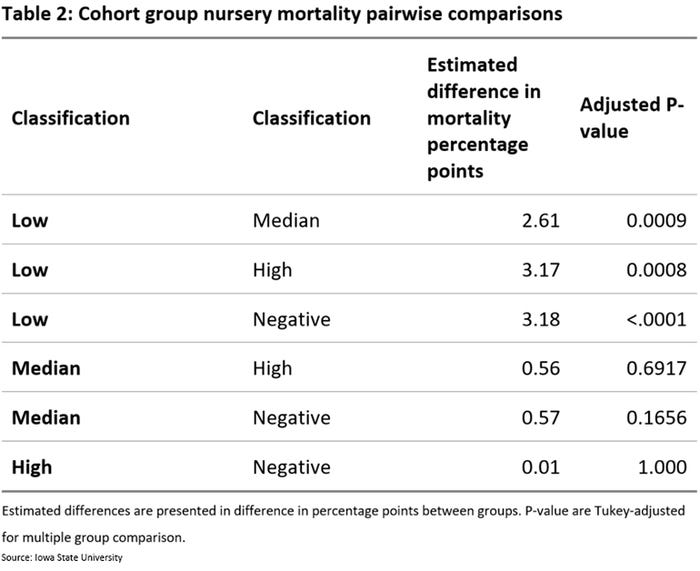
These results demonstrate that PRRS PCR Ct values results from processing fluids can be used to classify piglet cohort/batches, which appear to be associated with nursery performance. Specifically, for cohort/batches having low Ct values obtained on rRT-PCR for PRRSV are expected to have higher mortality than cohorts having median, high or negative results. One potential application of this would be to avoid mixing cohort group of piglets with low Ct values obtained on rRT-PCR for PRRSV with piglets of different Ct values when commingling for nursery placement.
References
1. Holtkamp D, Kliebenstein J, Neumann E, Zimmerman J, Rotto H, Yoder T, et al. Assessment of the economic impact of porcine reproductive and respiratory syndrome virus on United States pork producers. Journal of Swine Health and Production. 2013;21(2):72-84. PubMed PMID: WOS:000315319600005.
2. Lopez W, Angulo J, Zimmerman J, Linhares D. Porcine reproductive and respiratory syndrome monitoring in breeding herds using processing fluids. Journal of Swine Health and Production. 2018;26(3):146-50. PubMed PMID: WOS:000430994300007.
3. Holtkamp D, Polson D, Torremorell M. Terminology for classifying swine herds by porcine reproductive and respiratory syndrome virus status. Journal of Swine Health and Production. 2011;19(1):44-56. PubMed PMID: WOS:000285776000008.
4. Trevisan G, Linhares L, Crim B, Dubey P, Schwartz K, Burrough E, et al., editors. Swine Disease Reporting System: real-time, aggregated results from major U.S. Veterinary diagnostic laboratories. James D McKean Swine Conference; 2018; Ames-IA.
5. Trevisan G. Swine Disease Reporting System. Phase 1 Porcine Reproductive and Respiratory Syndrome Virus - PRRSv. available at Powerbi.com username: [email protected] password: Bacon 100 APP PRRSv ed2018.
You May Also Like



