An acute outbreak of atypical neurologic disease in 11-week-old pigs in a finishing barn was recently investigated by a group of veterinary diagnostic labs (Iowa State University, Kansas State University and the University of Minnesota).
October 3, 2016

An acute outbreak of atypical neurologic disease in 11-week-old pigs in a finishing barn was recently investigated by a group of veterinary diagnostic labs (Iowa State University, Kansas State University and the University of Minnesota). This was a collaborative effort between the authors as well as Kent J. Schwartz (ISU), Fabio Vannucci (UMN), Talita Resende (UMN), Albert Rovira (UMN), Paul Sundberg (Swine Health Information Center), Jerome Nietfeld (KSU), and Ben M. Hause (KSU).
Clinically affected animals originated from a single nursery and were placed in two different finishers two weeks prior to the initiation of clinical signs. Over the course of a three-week time frame, the onset and progression of clinical signs included decrease of water and feed consumption, compromised ambulation with ataxia (difficulty in walking), incoordination of limbs, mental dullness, paresis (muscular weakness), paralysis (unable to move affected limb or limbs) and decreased response to environmental stimuli. More severely affected animals had severe ataxia with intact deep pain perception and withdrawal reflexes in the hind limbs.
Despite mental dullness and aimless wandering, pigs often had central awareness with no detected nystagmus or cranial nerve deficits. Video of the affected pigs is available at swinehealth.org/sapelovirus. Overall morbidity (percentage of clinically affected animals) of 20% and case fatality rate (percentage of animals who showed clinical signs that died) of 30% were reported by the herd veterinarian. Samples were collected and submitted to the ISU-VDL by the herd veterinarian.
Microscopic examination of the brain revealed severe lymphoplasmacytic and necrotizing encephalomyelitis with multifocal areas of gliosis and neuron satelliosis suggestive of a neurotropic (brain) viral infection (Figure 1). Bacterial isolation attempts from multiple tissues from affected pigs did not demonstrate any bacterial pathogens that could explain these microscopic lesions.
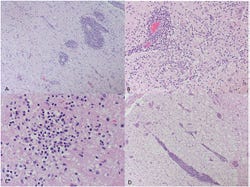
Figure 1: Upper left - Severe, multifocal lymphoplasmacytic encephalitis and gliosis. Upper right - Marked expansion of Virchow-Robin spaces by moderate to high numbers of lymphocytes and plasma cells. Locally extensive gliosis and neuron satelliosis. Lower left - Multifocal glial nodules with occasional neurophagia and satelliosis. Lower right - Multifocal, severe, lymphoplasmacytic myelitis and occasional spongiosis.
Immunohistochemistry and polymerase chain reaction testing for porcine reproductive and respiratory syndrome virus and porcine circovirus type 2 were negative. Sections of spinal cord tested by multiplex PCR for porcine enterovirus, porcine teschovirus and porcine sapelovirus were found consistently positive for only porcine sapelovirus A.
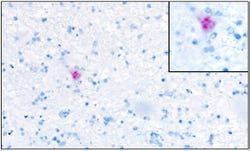
Figure 2: Recently developed Sapelovirus A in situ hybridization. Histologic section of cerebrum from affected pig. Neuron with intracytoplasmic positive staining. Figure on right corner shows a higher magnification of cell with positive staining.
Next-generation sequencing of brainstem and spinal cord samples was performed to investigate the possible presence of other infectious agents as well as to confirm previous sapelovirus PCR results. Interestingly, a genetically novel sapelovirus was identified within CNS tissues of affected animals.1 The 2,323 amino acid polyprotein sequence detected has an overall 94% amino acid identity and 86% nucleotide identity to a recently reported Korean sapelovirus2. No other viral agents were identified within examined samples, using the next generation sequencing. In order to further investigate the role of sapelovirus, in situ hybridization assay for porcine sapelovirus was developed through collaboration with the University of Minnesota VDL. Results from this assay demonstrated positive staining within glial fibrillary acidic protein positive cells and neuron from tissues of affected animals (Figure 2).
Porcine sapelovirus A is a non-enveloped virus with positive sense, single stranded RNA genome belonging to the Picornaviridae family, which has been re-classified into the three distinct genera (Enterovirus, Teschovirus and Sapelovirus). Historical seroprevalence reports suggest that a variety of enteroviruses are endemic within most commercial swine farms. Between Jan. 1, 2014, and June 21, 2016, a total of 65 cases were submitted to ISU-VDL for neurologic disease investigation which also had histopathology examination of neurologic tissue and nested PCR for PSV, PTV and PEV performed.
Thirty of these 65 cases had histologic neurologic lesions attributed to a viral infection. One or more viruses were detected in 19 of those 30 cases. PSV was detected in 14 of the 19 cases, either alone (six cases) or with PTV or PEV (four and three cases, respectively). All three viruses were detected in one case. The biologic relevance of this finding is not clear at this point as there is a significant gap of knowledge concerning the ecology, pathophysiology and the potential role of Sapelovirus A in cases of encephalomyelitis in swine. Also, detection of a virus of enteric origin such as Sapelovirus should be interpreted with caution as fecal contamination of samples is common.
Clearly, additional studies are warranted for improving understanding of the ecology, epidemiology and pathophysiology of porcine enteroviruses generally and Sapelovirus A specifically. To date, sapeloviruses detected in other species have not been reported to be associated with nervous disease. In the case reported here, a novel sapelovirus was the only agent detected associated with a unique clinical presentation of neurologic disease. At least one previous case report documents the neuroinvasive potential of porcine sapelovirus in swine3. A direct causal link between PSV and encephalomyelitis in swine remains to be proven and significant knowledge gaps in epidemiology, pathogenesis and biologic relevance of this potential pathogen remain. The Swine Health Information Center has created guidelines for identification and reporting of CNS cases and more information can be found at swinehealth.org/neurologic-syndromes.
References
1. Paulo Arruda, Kent Schwartz, Bailey Arruda, Albert Rovira, Jerome Nietfeld, Paul Sundberg, Ben Hause. Identification of a divergent strain of Sapelovirus associated with a severe polioencephalomyelitis outbreak in the United States. www.swinehealth.org/sapelovirus
2. Son, K.Y., Kim, D.S., Kwon, J., Choi, J.S., Kang, M.I., Belsham, G.J., Cho, K.O. 2014: Full-length genomic analysis of Korean porcine Sapelovirus strains. PLos One. 9, e107860
3. Schock A., Gurrala R., Fuller H., Foyle L., Dauber M., Martelli F., Scholes S., Roberts L., Steinbach F., Dastjerdi A. (2014) Investigation into an outbreak of encephalomyelitis caused by a neuroinvasive porcine sapelovirus in the United Kingdom. Vet Microbiol 172, 381-389.
You May Also Like



