Blueprint Issue: The pig’s gut microbiome plays a critical role in providing a protective intestinal barrier, digesting and metabolizing nutrients, anddeveloping as well as maintaining immunity.
April 9, 2018

By Megan C. Niederwerder, DVM, Ph.D., Kansas State University Department of Diagnostic Medicine and Pathobiology
The gut microbiome is the term used to describe the collection of microorganisms living in the gastrointestinal tract. These microorganisms are diverse and have been reported to equal or outnumber the cells of the host.
Also, the gut microbiome plays a critical role in providing a protective intestinal barrier, digesting and metabolizing nutrients, and developing as well as maintaining immunity. The birth route, diet, environment, infection and genetics all play important roles in initial microbial colonization of the piglet gut, as well as the continued shaping of the microbiome during the first several weeks of life1.
Role in combating infection
The relationship, balance and mechanistic interactions between these microbes in the gut is extremely complex and not well understood in states of health or disease. However, there is growing evidence indicating the beneficial role that gut microbiome diversity plays in the outcome of infectious diseases.
In humans and pigs, the impact of the microbiome on infectious disease has historically focused on enteric infections causing diarrhea, due to the seemingly inherent interactions of beneficial and pathogenic microbes in the gut. For example, pigs characterized as susceptible to enterotoxigenic Escherichia coli had a reduction in intestinal bacterial diversity compared to those pigs considered non-susceptible2.
More recently, the interest in the microbiome and its effects on infectious disease have expanded, to include pathogens primarily affecting the respiratory tract1.
As infectious respiratory disease is a major cause of morbidity and mortality in both livestock and humans, the gut microbiome has emerged as an alternative strategy in the control and prevention of these infections.
The gut-lung axis is the term used to describe how the gut microbiome — likely through products of microbial metabolism and modulation of systemic immunity — communicates with the respiratory tract and has the potential to affect the outcome following respiratory infections.
Several published studies have shown a beneficial association between microbiome diversity and outcome following infection with viral, bacterial and even fungal respiratory pathogens. For example, in mice models of human diseases, increased gut microbial diversity has been associated with reduced mortality and decreased lung lesions after respiratory infection with Streptococcus pneumoniae3.
Similarly, in pigs, increased gut microbial diversity has been associated with reduced clinical signs and decreased lung lesions after respiratory infection with Mycoplasma hyopneumoniae4.
Porcine reproductive and respiratory syndrome virus and porcine circovirus type 2 are two of the most important pathogens affecting the swine industry worldwide.
Co-infections are common on a global scale, resulting in pork production losses through reducing weight gain and causing respiratory disease in growing pigs. Both viruses modulate the immune system, increasing the risk of primary and secondary pathogens, and both viruses cause systemic infections with primary lung pathology.
Economic loss due to disease
Since PRRSV was introduced nearly three decades ago, it has cost an estimated $14 billion in losses to swine production. The most recent analysis of the cost of PRRS to the U.S. alone is $664 million yearly5.
In large part, these economic losses are associated with increased morbidity and mortality due to respiratory disease in growing pigs. PRRS-associated respiratory disease in the growing pig population not only reduces weight gain, but also increases secondary bacterial infections, increasing the need for antimicrobial treatments.
The currently available PRRS vaccines are generally considered inadequate for disease control, and programs designed for long-term elimination of PRRSV from a herd are often unsuccessful. The gut microbiome offers an alternative approach to improve weight gain and reduce morbidity and mortality associated with respiratory disease in the presence of PRRS.
Considering the potential impact that the gut microbiome has on respiratory pathogens, our work has focused on identifying associations between microbiome diversity and disease outcome characteristics in pigs, such as the presence of clinical signs and reduced growth, using a PRRSV-PCV2 co-infection model.
Put to the test
Our initial work demonstrated that the fecal microbiome was associated with clinical outcome of pigs 70 days post-infection with PRRSV and PCV26.
Specifically, reduced clinical signs, improved weight gain and reduced pathology was associated with increased microbiome diversity and the presence of Escherichia coli in feces. However, it remained uncertain if microbiome characteristics could predispose response to co-infection.
The purpose of the study described herein was to determine if microbiome characteristics present at the time of virus exposure were associated with outcome after co-infection with PRRSV and PCV27.
For this study, 3-week-old barrows (n = 50) were obtained at weaning from a high health commercial source negative for PRRSV. Pigs were given approximately four weeks to acclimate to their new environment prior to co-infection with PRRSV and PCV2.
Fecal samples were collected from all 50 pigs during the week prior to co-infection. During the 42-day post-infection period, body weights were collected weekly and blood samples were collected weekly to biweekly.
At 35 days post-infection, 20 pigs were selected to represent high-growth-rate pigs (n = 10) and low-growth-rate pigs (n = 10). These two groups were selected based on average daily gain post-infection, with pigs in the high-growth-rate group having the highest ADG and pigs in the low-growth-rate group having the lowest ADG.
At the conclusion of the trial, the 20 pigs selected had complete necropsies performed, and pathology was assessed in both the lung and lymphoid tissues.
Lung tissues were scored with regards to evidence and severity of interstitial pneumonia, and lymphoid tissues were scored with regards to lymphoid depletion, a characteristic finding in pigs with porcine circovirus-associated disease.
PRRSV and PCV2 viremia were compared between the two groups using quantitative PCR. Utilizing the Lawrence Livermore Microbial Detection Array, we profiled the microbiome in feces collected prior to co-infection from pigs in the high- and low-growth-rate groups.
The LLMDA provides the opportunity to detect over 10,000 microbes in a single test, including viruses, bacteria, fungi, protozoa and archaea species.
After co-infection with PRRSV and PCV2, significant variation occurs in a population of pigs, including extremes of weight gain and clinical disease (Figure 1)7.
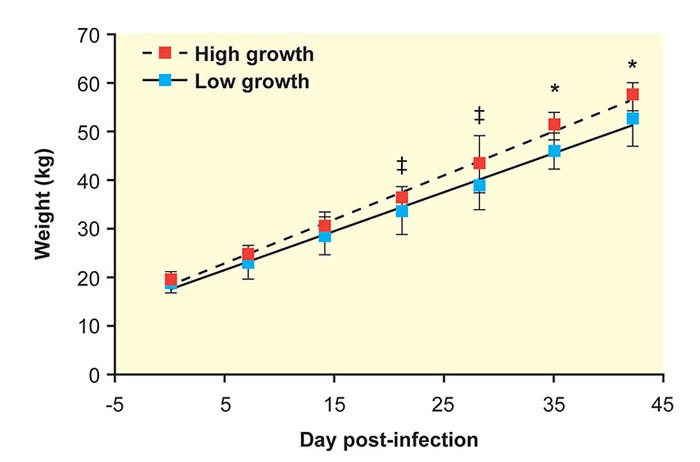
Figure 1: After co-infection with PRRSV and PCV2, significant variation occurs in a population of pigs. (Ober et al. 2017, Vet. Microbiol.)
Absolute weights of the high- and low-growth-rate pigs were similar at the time of co-infection but diverged after exposure to PRRSV and PCV2. Mean ADG for the 10 high-growth-rate pigs was 2 pounds compared to 1.7 pounds in the low-growth-rate group during the post-infection period.
Weight leader
At the conclusion of the study, the high-growth-rate group was 10.1 pounds heavier on average when compared to the low-growth-rate group.
When comparing virus replication between the two groups, high-growth-rate pigs had significantly less PRRSV and PCV2 detected in the serum, supporting the role of pathogen load in determining outcome during infection. Also, high-growth-rate pigs had less severe interstitial pneumonia.
Because the microbiome analysis was performed on pre-infection feces from pigs identified retrospectively as having high or low growth rates after co-infection, any effects of infection on the microbiome were excluded, and characteristics could be associated with predisposing or predicting outcome.
Overall, 29 microbial families and 112 individual microbial species were detected by the LLMDA in feces, including viruses, archaea and bacteria. At the level of the fecal microbiome, high-growth-rate pigs had increased microbial diversity on both a family and species level (Figure 2)7.
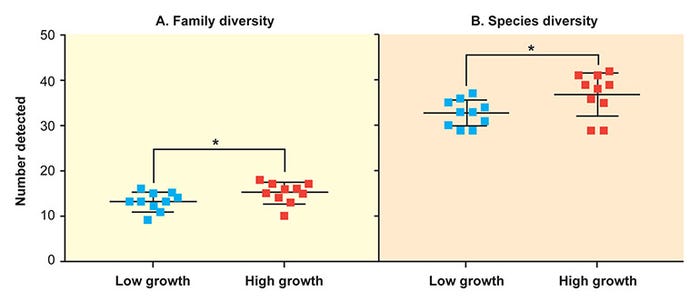
Figure 2: At the level of the fecal microbiome, high-growth-rate pigs had increased microbial diversity on both a family and species level. (Ober et al. 2017, Vet. Microbiol.)
The mean number of families in the low-growth-rate group was 13.1 compared to a mean of 15.1 in the high-growth-rate group. Similarly, the mean number of species in the low-growth-rate group was 32.7 compared to a mean of 36.9 in the high-growth-rate group.
Shifts in the microbiome composition of high-growth-rate pigs included reduced Methanobacteriaceae, increased Ruminococcaceae and increased Streptococcaceae species when compared to low-growth-rate pigs.
When taken together, these results indicate that both microbiome diversity and composition at the time of virus exposure may play a role in the subsequent response of pigs to PRRSV-PCV2 co-infection.
Profiling the microbiome characteristics in feces before infection provided a unique opportunity to consider possible avenues for prophylactic microbiome modulation as a preventative tool.
Oftentimes in studies published in the species of interest, microbiome associations with disease are characterized after the disease has already occurred, limiting our ability to understand how the microbiome may predispose the development of the disease or exclude the effect of the disease on the microbiome.
For example, several studies have characterized the microbiome of obese humans as having an enhanced capability to extract energy from food due to the relative abundance of two primary bacterial phyla, Firmicutes and Bacteroidetes8,9. However, these microbiome characteristics are typically defined in obese patients already exhibiting the disease condition.
In the current experimental infection study, we retrospectively analyzed the microbiomes of pigs before infection, allowing us to make associations between microbiome characteristics and subsequent outcome.
The role of the microbiome in PRRSV-PCV2 co-infection.
Altering microbiome
Beneficial characteristics of the pre-infection microbiome included increased diversity, as well as shifts in the presence of several bacterial families. Knowing that pre-infection associations exist suggests it may be possible to modulate or shift the microbiome to improve outcome during pathogen exposure.
This is a first step to understanding how the microbiome may be utilized as an alternative tool in swine for disease control and improving weight gain in the presence of infectious pathogens.
Alternatives to antimicrobials and alternative tools for disease control are an essential need for modern-day pork producers and the swine industry.
There is an urgent need for creative outside-the-box solutions to improve growth and health in swine while minimizing or eliminating the use of antimicrobials. This is particularly true for control of PRRS, as it continues to be the most costly disease of swine production, despite several decades of research.
The microbiome is a promising alternative strategy, due to its role in both weight gain and immunity, to promote health, improve response to infection and increase growth in swine. In addition, the gut microbiome has the advantage of being modulated orally, enabling a non-invasive administration route.
In the aforementioned study, the high-growth pigs with increased microbiome diversity weighed an average of 10.1 pounds more than the low-growth pigs after co-infection.
If these microbiome characteristics translated to increased weight gain in a PRRS-positive herd, the potential cost benefits could be significant.
Connection to human health
Modern-day swine production can draw interesting parallels to the hygiene hypothesis in humans, where increased sanitation, reduced exposure to microbes in early life and frequent use of antimicrobials have been associated with increased susceptibility to certain disease conditions1.
The overarching theme of the current work is that increased microbial diversity is associated with improved outcome following infectious pathogen exposure.
It begs the question: When broadly treating swine herds with antimicrobials, could we be reducing microbial diversity in the gut and harming beneficial microbes?
Overall, it is important to remember that the gut microbiome and overall gut health has the potential to impact the health of the whole pig, including the respiratory tract in response to diseases such as PRRS.
As we move forward with reducing antimicrobial usage and increased antimicrobial stewardship in swine production, much is to be learned about how we best “take care of” beneficial microbes to improve the response of pigs to infectious diseases.
References
1. Niederwerder, M.C., “Role of the microbiome in swine respiratory disease.” Vet Microbiol, 2017. 209: p. 97-106.
2. Messori, S., et al., “Effect of susceptibility to enterotoxigenic Escherichia coli F4 and of dietary tryptophan on gut microbiota diversity observed in healthy young pigs.” Vet Microbiol, 2013. 162(1): p. 173-9.
3. Schuijt, T.J., et al., “The gut microbiota plays a protective role in the host defense against pneumococcal pneumonia.” Gut, 2016. 65(4): p. 575-83.
4. Schachtschneider, K.M., et al., “Modulation of systemic immune responses through commensal gastrointestinal microbiota.” PLoS One, 2013. 8(1): p. e53969.
5. Holtkamp, D.J., et al., “Assessment of the economic impact of porcine reproductive and respiratory syndrome virus on United States pork producers.” Journal of Swine Health and Production, 2013. 21(2): p. 72-84.
6. Niederwerder, M.C., et al., “Microbiome associations in pigs with the best and worst clinical outcomes following co-infection with porcine reproductive and respiratory syndrome virus and porcine circovirus type 2.” Veterinary Microbiology, 2016. 188: p. 1-11.
7. Ober, R.A., et al., “Increased microbiome diversity at the time of infection is associated with improved growth rates of pigs after co-infection with porcine reproductive and respiratory syndrome virus and porcine circovirus type 2.” Vet Microbiol, 2017. 208: p. 203-211.
8. Turnbaugh, P.J., et al., “An obesity-associated gut microbiome with increased capacity for energy harvest.” Nature, 2006. 444(7122): p. 1027-31.
9. Ley, R.E., et al., “Microbial ecology: human gut microbes associated with obesity.” Nature, 2006. 444(7122): p. 1,022-3.
You May Also Like



