August 31, 2015
Back in high school, 25 years ago, in our philosophy class, the professor introduced the concept, “No man ever steps in the same river twice,” which is attributed to the Greek philosopher Heraclitus.
Admittedly, I had to search for the quote to remember the name of the philosopher. However, those of us who were born and raised next to a river understand what the ancient philosopher meant. In a sense, there are certain features that are constant in a given river. Its name, source, course and mouth are always the same. Yet, we never step in the same river twice. There are days in which the river is placid and serene, and there are days in which it is furious and energetic. The river has certain features that are constant, but it is never the same.
Something similar occurs with RNA viruses, like the porcine reproductive and respiratory virus. There are certain features that are constant in the PRRSV, most notably, its structure and conformation. PRRSV is, however, not always the same. The virus is able to infect pig cells thanks to some specific proteins, and the immune system of the animals tries to prevent their action by developing specific protection against those proteins.
The structure of those proteins is determined by the genetic material of the virus, whose minimum units are called “bases.” During the virus replication process, when new viruses are generated, there are “mistakes” in the 15,000 base-long string of genetic material, and some of the bases may change. As a result, the virus is still a PPRSV, but like the river, the virus proteins are now slightly different. Also, as the river has calm and furious days, there are variants of the PRRSV that are more aggressive than others.
Aggressive PRRS variant
Recently, an aggressive variant of the PRRSV, with a 1-7-4 RFLP pattern, was responsible for causing some particularly harmful outbreaks, with evident clinical manifestation of disease throughout the country. The bug rapidly spread through some farms and systems resembling, again, what in a river would be a “storm.”
Does it mean that the PRRSV will now be more clinically evident than in the past? We do not think so. The immune system of the animals will develop protection, and like in the river, this storm will pass and there will be quieter days. However, it is not until we can dry the river (or eliminate the disease) that storms will stop harming us.
A recent project at the University of Minnesota is intended to analyze the changes in the genetic composition of the PRRSV to be able to identify the routes through which the virus spread. Analyzing the genetic structure of the virus from samples collected in the outbreaks, we can know how bad is the current rate of virus mutation, when compared to earlier outbreaks, and reconstruct the most likely history of spread between and among farms. This information will help to identify the routes of spread and, most important, with time, we expect to develop a system that could serve as an alarm, like a bad-weather alert, that producers and veterinarians may use to anticipate when conditions are favorable for storms to occur.
Half of production sows share data
A number of participants have joined efforts to share data and information, on a confidential basis, to build this system, through the Swine Health Monitoring Program, that includes approximately half of the population of sows in the United States. As we progress in our research, we expect that time will come in which, like we check the weather forecasting every morning, we will be able to check this system of alerts for potentially harmful PRRSV strains. The storm may still come, but we will be better prepared.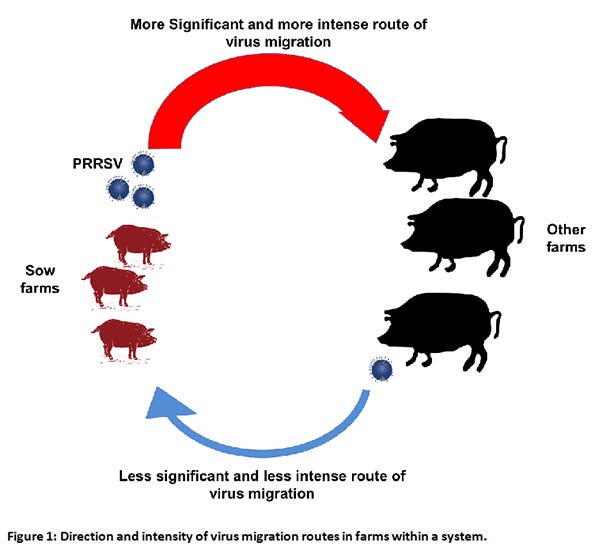
So to test the biological soundness (i.e. IQ) of our sequences analysis model, we basically asked the following questions - given our collected sequence data within a system:
What is the most likely origin of this new 1-7-4 strain, would it mostly likely have come from a sow farm or another type of farm (finishing, GDU, etc…)?;
What is the direction of spread between both types of farms?;
How intense is the movement of the genetic material of the virus between these farms?
The model’s answers were (see Figure 1):
1-7-4 strain most likely originated from sow farms
The direction of spread is strongly significant from sow farms to other farms (color red);
More viruses are moving from sow farms to other farms (Thicker arrow).
So, what should we do with this information? We might want to revise biosecurity measures in our farms, review animal and people movements and intensify sampling efforts in sow farms.
You May Also Like



