The development of these antibody tests will soon give veterinarians another tool in SVA surveillance.
September 18, 2017
By Russ Daly and Steve Lawson, South Dakota State University
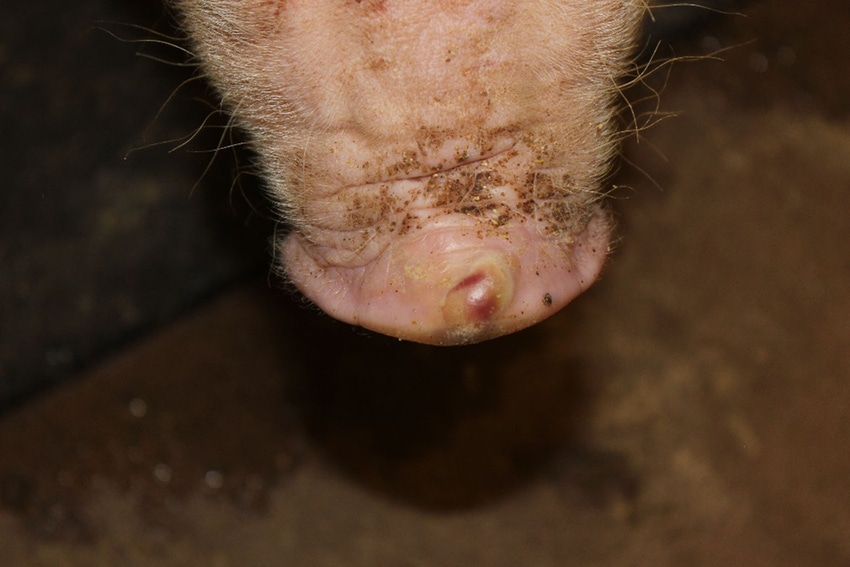
Clinical problems due to Senecavirus A, formerly known as Seneca Valley Virus, have quickly expanded from sporadic oddities in far-flung groups of pigs to well-established infections throughout the pig-producing world. The virus is implicated in outbreaks of vesicular (blister-like) sores on snouts and feet of growing pigs, as well as increased mortality in baby pigs.
While SVA can cause these production losses, it’s perhaps even more problematic because of how its clinical signs are identical to foot-and-mouth disease. When vesicles pop up in pig populations, they need to be investigated with the urgency of a foreign animal disease. This makes diagnosis of SVA — and its proper differentiation from FMD — a critical need for regulatory and clinical veterinarians.
The veterinary diagnostic community, including the South Dakota Animal Disease Research and Diagnostic Laboratory, quickly responded to the emergence of SVA by developing molecular-based polymerase chain reaction testing to detect the virus in tissue samples and swabs from affected pigs. This test has been widely adopted by veterinary diagnostic laboratories and has greatly aided veterinarians in sorting out vesicular problems in pigs.
The PCR testing process, however, can be involved and relatively expensive, and depends upon the immediate presence of the virus (or its nucleic acid) in the tested population. What if there were diagnostic methods for SVA that were less expensive and could detect ongoing — maybe even subclinical — exposures, therefore better lending themselves to use for routine and regular surveillance?
That’s where serology comes into play — the detection of antibodies in the animal’s bloodstream, indicating previous exposure to a disease agent. Serologic tests for SVA are rapidly being developed, and much of that work is being done by researchers at the SD ADRDL, including a team consisting of Steve Lawson, Eric Nelson, Diego Diel, Aaron Singrey, Travis Clement and Jane Hennings.
By definition, serologic tests indicate the presence of antibodies in a sample by taking advantage of the very specific, very strong bond between the antibody and an antigen (a germ, or more typically, a protein molecule specific to that germ). This interaction then has to “made visual,” often through the addition of fluorescent markers to the test components.
The work that goes into developing an accurate serologic test cannot be overestimated, especially for emerging pathogens such as SVA.
Researchers must first work with the virus itself, in order to find the specific portions, often structural proteins, of the virus that are the best “targets” for antibodies. In this case, SDSU researchers took the whole genome sequence of a typical SVA strain and compared it to the sequences of different picornaviruses (the family containing SVA). This step ensured that the SVA test would only identify SVA and not other closely related viruses. After these SVA-specific proteins are identified, they’re “manufactured” by cloning specific nucleic acid from the SVA into E. coli vectors, which produce the proteins in large enough quantities to use in test development.
Then it falls to a lot of trial-and-error to see which of these protein candidates work the best in a test to detect antibodies. Useful in this process are real-world serum samples from pigs that have gone through SVA outbreaks, as well as from pigs on farms that have never seen the virus. In the case of the SDSU work, a structural protein known as VP2 became the best candidate to use in serology.
The SDSU group is working on several different serologic tests, including virus neutralization, indirect fluorescent antibody, competitive ELISA, fluorescent microsphere immunoassay, as well as others. Most of these use the purified SVA proteins in various arrangements with antibodies and fluorescent markers or enzymes to visualize antibody in a sample. All of these tests have their own plusses and minuses, with the virus neutralization tests in general showing the best specificity (fewer false positives) and the ability to detect an SVA-positive animal in as little as five to seven days post-infection. Work continues on refining the other methods, particularly the ELISA tests that can be more efficiently automated.
The development of these antibody tests will soon give veterinarians another tool in SVA surveillance. Serologic testing is also critical for manufacturers testing SVA vaccine candidates, and offshoots of the test development process such as monoclonal antibodies are important in diagnostic processes such as virus isolation. As time goes on, researchers continue to refine these tests to make them more and more applicable to all of us trying to sort out this confounding virus.
You May Also Like



